People
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0003-1613-4663
https://orcid.org/0000-0003-1613-4663
Projects: RootBook
Institutions: University of Oslo
Projects: Auromega, AquaHealth (ERA-BlueBio)
Institutions: SINTEF
Projects: HubMOL
Institutions: UiT The Arctic University of Norway
Project manager HubMOL
Projects: COVID-19 Disease Map
Institutions: University of Cape Town
 https://orcid.org/0000-0002-5374-0657
https://orcid.org/0000-0002-5374-0657
Projects: BullNet
Institutions: University of Cologne
Projects: SilicoTryp
Institutions: University of Glasgow
Projects: COVID-19 Disease Map
Institutions: University of Pittsburgh
Projects: INDIE - Biotechnological production of sustainable indole
Institutions: Axxence Aromatic GmbH
Project leader at Axxence Aromatic
Professor in Neuroscience and Molecular Pharmacology at UMCU, Utrecht, NL
Projects: Not specified
Institutions: Not specified
Customized AE is a trusted provider dedicated to delivering high-quality, one-stop gifting solutions tailored to your needs. Our expertise covers a wide range of personalized products, including customized mugs near me, creative designs, and premium printing services to help individuals and businesses make a lasting impression. Whether you are searching for mug printing near me or unique custom cups dubai, we ensure reliable service, durable quality, and ...
Projects: Xenophiles Systems Biology
Institutions: VU University Amsterdam
Ph D student of Hans Westerhoff at VU working on Xeonomicrobiology
Projects: Xenophiles Systems Biology
Institutions: VU University Amsterdam
Projects: SysVirDrug
Institutions: NovaMechanics Ltd
Projects: Not specified
Institutions: Not specified
"BalaJi MicroTechnologies BMT is New Delhi, india based company. We are ISO 9001:2015 Certified company & registered in India under Indian companies act. We are a Unit of ""B.B. Group of Companies"".
""India's Biggest Complete Industrial Automation Solution Supplier""
BMT is India's biggest ""System Integrator & Distributor"" for several Industrial Automation Applications in Packaging, Pick & Place, Printing, Control Panels, Textile, Pharma, FMCG, Bottling, Vision Inspection machines, ...
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: HubMOL
Institutions: Novo Nordisk Foundation Center for Basic Metabolic Research
 https://orcid.org/0009-0001-3213-7285
https://orcid.org/0009-0001-3213-7285
Projects: COVID-19 Disease Map
Institutions: Universität Konstanz
 https://orcid.org/0000-0002-1286-1372
https://orcid.org/0000-0002-1286-1372
Projects: Compound Data, BioTarget Database
Institutions: Università degli Studi di Modena e Reggio Emilia (UNIMORE)
Projects: ZucAt, Modelling COVID-19 epidemics
Institutions: Institute of Cytology and Genetics, Novosibirsk State University
I am staff scientist in the lab of molecular-genetic systems at the Department of Systems Biology, Institute of Cytology and Genetics SB RAS and Postdoc Research Fellow at San Diego State University. My research focus is dynamical modeling of gene network functioining.
Projects: POP - the Parameter Optimisation Problem
Institutions: University of Exeter
 https://orcid.org/0000-0002-9320-5958
https://orcid.org/0000-0002-9320-5958
Projects: Smart Garden Watering System
Institutions: Tech Mahindra
Projects: STREAM
Institutions: University of Groningen
Mathematical modeling, bioinformatics
Projects: SteaPKMod, Quantifizierung des Zusammenhanges zwischen Leberperfusion und -funktion bei erweiterter Leberresektion - Ein systemmedizinischer Ansatz (QuaLiPerF)
Institutions: Experimental Transplantation Surgery, Department of General, Visceral and Vascular Surgery, University Hospital Jena, Jena University Hospital
 https://orcid.org/0000-0002-6192-8713
https://orcid.org/0000-0002-6192-8713
Projects: SulfoSys - Biotec
Institutions: University Bielefeld
Projects: SulfoSys, SulfoSys - Biotec
Institutions: Max-Planck-Institute for terrestrial Microbiology
Projects: SulfoSys - Biotec
Institutions: University Bielefeld
 https://orcid.org/0000-0002-3394-1304
https://orcid.org/0000-0002-3394-1304
Projects: MESI-STRAT
Institutions: Medical University Innsbruck
Projects: SysMO DB
Institutions: University of Manchester - Department of Computer Science
Intern @ myGrid team
Projects: Glycogen Metabolism in bacteria, Diverse Metabolic Control of Phosphoglucomutases by Bisphosphorylated Sugars in Heterotrophic Bacteria, Controlled Carbon Loss: Threshold-Dependent Overflow Metabolism in Synechocystis
Institutions: University of Tuebingen
 https://orcid.org/0009-0009-1115-5175
https://orcid.org/0009-0009-1115-5175
Projects: WineSys, INBioPharm, BioZEment 2.0, CoolWine, Auromega
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-9125-326X
https://orcid.org/0000-0002-9125-326X
Projects: KOSMOBAC
Institutions: University of Aberdeen
Projects: Hydrocow
Institutions: RWTH Aachen University - Department ABBt, iAMB
 https://orcid.org/0000-0002-9593-4388
https://orcid.org/0000-0002-9593-4388
Projects: Cross-Feeding B.theta
Institutions: University of Würzburg
 https://orcid.org/0000-0002-5066-7140
https://orcid.org/0000-0002-5066-7140
Projects: Environmentally-friendly bioadhesives from renewable resources (WooBAdh)
Institutions: Université de Lorraine
Projects: FAIRDOM Community Workers, FAIRDOM, Cell4Chem
Institutions: University of Manchester - Department of Computer Science, University of Manchester
Projects: SysMilk
Institutions: European Molecular Biology Laboratory
 https://orcid.org/0000-0002-7875-0261
https://orcid.org/0000-0002-7875-0261
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: University of Freiburg
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-4056-0606
https://orcid.org/0000-0002-4056-0606
Projects: EmPowerPutida
Institutions: University of Stuttgart
Projects: The Beer Project
Institutions: Norwegian University of Science and Technology
Projects: MESI-STRAT
Institutions: PATH - Patients' Tumor Bank of Hope
Projects: CoolWine
Institutions: Universitat Rovira i Virgili
Projects: COVID-19 Disease Map
Institutions: Shanghai University
Projects: FAIRDOM
Institutions: University of Manchester - Department of Computer Science
Projects: FAIRDOM user meeting
Institutions: CEIT
Projects: Pan-cancer multiSOM project
Institutions: Institute of Molecular Biology NAS RA
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: CC-TOP
Institutions: Université Clermont Auvergne (UCA)
Projects: CC-TOP
Institutions: CSIC Barcelona - Consejo Superior de Investigaciones Científicas
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: FAIRDOM user meeting
Institutions: IDIBAPS
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: COVID-19 Disease Map
Institutions: King AbdulAziz University
Projects: SysMO-LAB
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
EISBM President - Disease Map Community Founder and Principal Investigator
Projects: COVID-19 Disease Map
Institutions: Sanofi R&D
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Projects: FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: University of Rostock
Projects: COVID-19 Disease Map
Institutions: Oregon Health & Science University (OHSU)
Projects: COVID-19 Disease Map
Institutions: Harvard Medical School
Projects: NGID
Institutions: Radboud University Medical Center
Projects: SUSPHIRE - Sustainable Bioproduction of Pheromones for Insect Pest Control in Agriculture, INDIE - Biotechnological production of sustainable indole, ADAPT - Accelerated Development of multiple-stress tolerAnt PoTato
Institutions: National Institute of Biology
 https://orcid.org/0000-0003-4776-7164
https://orcid.org/0000-0003-4776-7164
I am a biochemist & bio-informatician working in phytobacteriology at the Plant Sciences Unit of ILVO, the Flanders Research Institute for Agricultural, Fisheries and Food Research. The focus is on genomics-based research and diagnostics for Plant Health.
My expertise is 'wet-lab' work (microbiology, sequencing, molecular biology, design & validation of diagnostics assays using qPCR/LAMP, automatisation) and 'dry-lab' work such as bio-informatics/data analysis (e.g. scripting analysis ...
Projects: Automated Hepatic Vessel Segmentation in Whole-Slide Images Using U-Net Variants with Attention and SE Blocks, Quantifizierung des Zusammenhanges zwischen Leberperfusion und -funktion bei erweiterter Leberresektion - Ein systemmedizinischer Ansatz (QuaLiPerF)
Institutions: Experimental Transplantation Surgery, Jena University Hospital
 https://orcid.org/0009-0001-7234-3905
https://orcid.org/0009-0001-7234-3905
Projects: TRANSLUCENT
Institutions: Humboldt-Universität zu Berlin
Projects: COSMIC
Institutions: University of Rostock
Prof. Dr. Hubert Bahl Head, Division of Microbiology, Institute of Biological Sciences University of Rostock Albert-Einstein-Str. 3 D-18051 Rostock
Projects: EmPowerPutida
Institutions: ETH Zurich
Projects: SUMO
Institutions: University of Sheffield
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
I am a Professor in Medical Systems Biology and the University Medical Centre Groningen. The research in my lab is focused on complex regulation of mammalian lipid and carbohydrate metabolism, eventually aiming at network-based therapies. We combine dynamic computer simulations with quantitative metabolomics, 13C fluxomics, proteomics and transcriptome analysis, and in depth biochemical analysis. This allows to predict and understand ‘emergent’ properties, those properties that are counterintuitive ...
Postdoctoral researcher at Luxembourg Centre For Systems Biomedicine (LCSB), University of Luxembourg
Projects: COVID-19 Disease Map
Institutions: University of Cambrige
Projects: MIX-UP
Institutions: RWTH Aachen University - Department ABBt, iAMB
 https://orcid.org/0000-0001-5729-1724
https://orcid.org/0000-0001-5729-1724
We are very interested in applying a systems approach (i.e. model-based concepts and related computational tools) to problems from the biological domain. In particular, we are doing research in computational systems biology, targetting the following topics:
- Parameter estimation (inverse problems, model calibration) in biochemical pathways
- Optimal experimental design (optimal dynamic experiments) for Systems Biology
- Dynamic optimization (optimal control) of biosystems and bioprocesses
...
Projects: COVID-19 Disease Map
Institutions: Shanghai University
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University Medical Centre Groningen
Projects: COVID-19 Disease Map
Institutions: University of Surrey
 https://orcid.org/0000-0001-5640-7422
https://orcid.org/0000-0001-5640-7422
Reader (Professor) of Systems Biology; Executive Director for the International Society of Systems Biology (ISSB); Editor-in-Chief of Current Opinion in Systems Biology (Elsevier).
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: MESI-STRAT
Institutions: University of Innsbruck
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: FAIRDOM
Institutions: ETH Zurich & Basel / The Scientific IT Services (SIS) division
Projects: CC-TOP
Institutions: University of Groningen
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: SilicoTryp
Institutions: University of Glasgow
Projects: SysVirDrug
Institutions: University of Heidelberg
Projects: BaCell-SysMO
Institutions: University of Erlangen-Nuernberg
Projects: NIRBTEST
Institutions: University Duisburg-Essen
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: COVID-19 Disease Map
Institutions: Biomax Informatics AG
Projects: WG Infrastructure for Translational Research, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", FAIRDOM user meeting, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University Medical Center Göttingen
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Projects: BESTER
Institutions: University of Ulm
Projects: BESTER
Institutions: University of Ulm
Projects: FAIRDOM
Institutions: Heidelberg Institute for Theoretical Studies
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: WURSynBio, INDIE - Biotechnological production of sustainable indole
Institutions: Wageningen University & Research
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: Core Facility Fluorescence Cytometry
Institutions: Forschungszentrum Borstel
Projects: iRhythmics
Institutions: University of Rostock
Institutions: University of Amsterdam
Martijn Bekker (1979) was born in Amstelveen (The Netherlands). He started his studies in biology in 1997 at the University of Amsterdam, and graduated in 2003 with specializations in molecular microbiology and in immunology. The internships during his undergraduate studies were carried out in the labs of Prof. dr. B. Oudega (VU, Amsterdam, The Netherlands) and Prof. dr. F. Heffron (OHSU, Portland, Oregon, USA). He continued with his graduate studies in 2003 in the Laboratory for Molecular Microbial ...
Projects: Not specified
Institutions: Not specified
https://amazonsale.io/ https://amazonsale.io/eportfolios/1/Contact/Contact https://amazonsale.io/eportfolios/10/Just_Keto_Diet_Amazon/Just_Keto_Diet_Amazon https://amazonsale.io/eportfolios/100/LeptoFix_Amazon/LeptoFix_Amazon https://amazonsale.io/eportfolios/1000/Where_To_Get_Keto_Extreme_Fat_Burner/Where_To_Get_Keto_Extreme_Fat_Burner https://amazonsale.io/eportfolios/1001/Keto_Extreme_Fat_Burner_New_Zealand/Keto_Extreme_Fat_Burner_New_Zealand ...
Projects: SysMetEx
Institutions: Ruhr University Bochum
Projects: SysMO-LAB
Institutions: VU University Amsterdam
Projects: CoolWine
Institutions: Universitat Rovira i Virgili
 https://orcid.org/ 0000-0002-7071-205X
https://orcid.org/ 0000-0002-7071-205X
Projects: COVID-19 Disease Map
Institutions: Harvard Medical School
 https://orcid.org/0000-0001-5934-966X
https://orcid.org/0000-0001-5934-966X
Projects: LEANPROT
Institutions: biotechrabbit GmbH
Projects: SYSTERACT
Institutions: University of Tübingen
Projects: Noisy-Strep
Institutions: University of Cologne
With a background in the statistical mechanics of disordered systems, my research focusses on the interface between statistical physics and molecular biology. At the centre is the relationship between fluctuations and noise in biological systems, the corresponding statistical ensembles, and biological function. This connection emerges at very different levels and timescales, from stochastic modeling of gene expression to the population dynamics of regulatory DNA.
Projects: MESI-STRAT
Institutions: Medical University Innsbruck
Projects: EmPowerPutida
Institutions: LifeGlimmer GmbH
Projects: NIRBTEST
Institutions: Institut Curie
Projects: FAIRDOM user meeting
Institutions: Unicamp - University of Campinas
Projects: Rhodolive
Institutions: Latvia University of Agriculture
I am interested in the coupling of global regulation and metabolism in E. coli. To analyze this I construct and analyze defined mutant strains. These strains are characterized in bioreactor experiments of different types (batch, conti, pulse ...) and measurements on the level of metabolites, mRNA, and protein are applied. For all projects there are cooperation partners that use the data in modeling approaches either from the MPI Magdeburg or from the SUMO consortium.
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: testproject
Institutions: University Medical Center Göttingen, Department of Medical Informatics
Projects: PSYSMO
Institutions: Queen's University Belfast
Projects: Not specified
Institutions: Not specified
Galaxy Education is pioneered as a tech-driven and top-notch platform for multitude of educational indulgence for the students that provides various educational supportive services and student development training programmes all over India and Abroad. Galaxy Education was formulated in the year 2004 as a unit of Genixo Info Solutions Pvt. Ltd. We were the numero uno for coming up with categorically focused education modules for the students to grab skills that ...
Projects: Noisy-Strep
Institutions: University of Cologne
I am a phd student working on statistical physics and complex systems and applying concepts from these fields to biology.
Projects: SDBV/HITS
Institutions: Heidelberg Institute for Theoretical Studies
Projects: SysVirDrug
Institutions: German Cancer Research Center (DKFZ)
 https://orcid.org/0000-0002-5805-6109
https://orcid.org/0000-0002-5805-6109
Projects: Not specified
Institutions: Not specified
Creative Biolabs provides professional custom service packages to fully meet client's specific needs in drug development journey, especially for various antibody related modalities (monoclonal antibody, single domain antibody, bispecific antibody, multi-specific antibody, ADC, CAR T, etc.).
Projects: Not specified
Institutions: Not specified
CD BioSciences, a pioneering company in the biotechnology sector, has recently unveiled its eco-friendly bioenvironmental microorganism products. CD BioSciences has been at the forefront of providing high-quality services in biotechnology and life sciences worldwide. With a focus on sustainability, the company is committed to supporting the transition toward eco-friendly solutions in various fields, including bioenergy, ...
Projects: SulfoSys
Institutions: University of Bergen
Projects: CC-TOP
Institutions: Johnson Matthey
Projects: KOSMOBAC
Institutions: University of Aberdeen
Projects: BullNet
Institutions: University College Dublin
Projects: PSYSMO
Institutions: University of Dortmund
Projects: SysMilk
Institutions: European Molecular Biology Laboratory
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0001-9489-1657
https://orcid.org/0000-0001-9489-1657
Projects: SynBio4Flav
Institutions: Consejo Superior de Investigaciones Científicas
Projects: HYp - Spatiotemporal analysis of hypersensitive response to Potato virus Y in potato, pISA-tree, INDIE - Biotechnological production of sustainable indole, _p_stRT, Public project, Playground, SUSPHIRE - Sustainable Bioproduction of Pheromones for Insect Pest Control in Agriculture
Institutions: National Institute of Biology, NewHorizon
 https://orcid.org/0000-0001-7484-6031
https://orcid.org/0000-0001-7484-6031
Projects: Methylation differences during respiratory infection
Institutions: German Sport University Cologne
Projects: ROBUSTYEAST
Institutions: Freie Universität Berlin
Projects: COVID-19 Disease Map
Institutions: Sanofi R&D
Projects: TRANSLUCENT
Institutions: University of Bonn
Projects: FAIRDOM user meeting
Institutions: ZonMw
Projects: Dynamic interplay between nuclei and mitochondria in ageing cells
Institutions: National Institute on Aging
Projects: GLASS-NL
Institutions: Amsterdam UMC
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: Mimetas
Projects: Cell4Chem
Institutions: Helmholtz Zentrum für Umweltforschung (UFZ)
 https://orcid.org/0000-0002-1934-7456
https://orcid.org/0000-0002-1934-7456
I am a Deputy Director of Vinogradsky Institute of Microbiology, Russian Academy of Sciences, Moscow, Russia and the head of the laboratory in this Institute. I ams working in the field of diversity, ecology and metabolism of thermophilic prokaryotes. Together with colleagues we've described many new taxa of thermophilic prokaryotes including those of high level (families, orders, levels). I m especially interested in isolation and description of thermophilic prokaryotes with unusual types of ...
Projects: MOSES
Institutions: VU University Amsterdam
Projects: KOSMOBAC
Institutions: University of Aberdeen
My research interests are in the physiology of bacteria subjected to stress. The focus of my recent research has been the structure and function of regulated transport systems and ion channels involved in cellular homeostasis. These transporters and channels respond to specific signals by a change in activity that either corrects the imposed stress or protects the cell during exposure to the stress. Our systems biology interests are in the interplay of different enzymes systems and transporters ...
Projects: DigiSal-BT8121
Institutions: NORCE Norwegian Research Centre AS
 https://orcid.org/0000-0002-6975-755X
https://orcid.org/0000-0002-6975-755X
My PhD project is part of the NFR project SLAM-DUNK (RCN Project number: 326861). The main aim of SLAM-DUNK is to design, optimize, and integrate a combination of novel technologies (anaerobic digestion, microwave assisted pyrolysis, and microalgae cultivation) for the conversion of fish sludge to valuable products. The PhD project is focusing on the optimization of microalgae cultivation on residual streams from aquaculture directly and from digester liquid from an anaerobic digestion (AD) ...
Projects: BioZEment 2.0
Institutions: Oslo and Akershus University College
Projects: HUMET Startup
Institutions: University of Gothenburg
 https://orcid.org/0000-0003-0786-8091
https://orcid.org/0000-0003-0786-8091
POSITION: Professor of Cardiovascular Research at University of Gothenburg, chair of Dept of Molecular and Clinical Medicine, Institute of Medicine, University of Gothenburg, and part-time physician at Clinical Chemistry, Sahlgrenska University Hospital, Gothenburg, Sweden
RESEARCH: My main long-term interest has been to understand the underlying mechanisms that lead to, and consequences of, lipid accumulation in the liver, arterial wall and heart, with the goal of translating this knowledge into ...
Projects: TRANSLUCENT
Institutions: Humboldt-Universität zu Berlin
Projects: SCaRAB, STREAM, SYSTERACT, INBioPharm
Institutions: SINTEF, Norwegian University of Science and Technology
Senior Research Scientist at SINTEF, Dept. of Biotechnology and Nanomedicine, Research Group Mass Spectrometry
Projects: COVID-19 Disease Map
Institutions: Universität Konstanz
Projects: COVID-19 Disease Map
Institutions: University Medical Center Hamburg - Eppendorf Hamburg
Projects: middle ear
Institutions: Technische Universität Dresden
 https://orcid.org/0000-0002-3061-0171
https://orcid.org/0000-0002-3061-0171
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: University of Freiburg
Projects: FAIRDOM user meeting
Institutions: SYSBIO - Centre of Systems Biology
 https://orcid.org/0000-0002-6566-2844
https://orcid.org/0000-0002-6566-2844
Projects: KOSMOBAC
Institutions: Max-Planck-Institute of Biochemistry Martinsried
I am a PhD student working the group of Zoya Ignatova. Cellular and extracellular changes like crowding and osmotic stress conditions play a major role in protein aggregation. A change in the cytoplasmic composition is the result of an interplay between high osmotic pressures outside the cell volume and the cellular response to it in terms of uptake of K+ and secondary organic osmolytes. My research focuses on elucidating the role of natural osmolytes (known also as chemical chaperones or compatible ...
Projects: INDIE - Biotechnological production of sustainable indole
Institutions: Wageningen University & Research
Research scientist in multi-Omics, molecular (cell)biology and bioinformatics.
Projects: COVID-19 Disease Map
Institutions: European Institute for Systems Biology and Medicine
Projects: WATCHER
Institutions: Universidade de Santiago de Compostela
Projects: Creative Biogene
Institutions: Creative Biogene
As a scientists-founded and scientists-driven leading biotechnological company, Creative Biogene focuses on developing high quality products and services, as well as proprietary technology to support the research in the field of basic life sciences research, biomedical development, industrial synthetic application and preclinical drug discovery.
Projects: HOTSOLUTE
Institutions: University Duisburg-Essen
Projects: BESTER
Institutions: PROCESSIUM
Projects: Lipofungi, DeepHyperSpec, Bio4Fuels, LIGNOLIPP
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6287-3964
https://orcid.org/0000-0001-6287-3964
Projects: Sustainable co-production
Institutions: Sunchem Italy
Projects: Pectobacterium pangenome
Institutions: Wageningen University & Research
 https://orcid.org/0000-0003-0536-7787
https://orcid.org/0000-0003-0536-7787
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München / Institute of Experimental Genetics
Projects: MetApp, LEANPROT, C1Pro
Institutions: SINTEF, Norwegian University of Science and Technology
Since 1st of January 2015 I am Professor in synthetic biology at the Norwegian University of Science and Technology (NTNU) institute of biotechnology. Before that I was research director in the non-profit research institution SINTEF. My major research activities are within microbial molecular biology, mainly combining metabolic engineering, synthetic biology and systems biology to develop microbial cell factories, and focusing both on the products and on the raw materials. The research includes ...
Projects: COSMIC
Institutions: University of Ulm
Phd student University of Ulm Institute of Microbiology and Biotechnology Albert-Einstein- Allee 11 89069 Ulm, Germany
Projects: STREAM, SilicoTryp
Institutions: University of Groningen
I am currently Professor of Systems Biology at the University of Manchester. My research interests focus on the development of innovative computational approaches for post-genomic systems biology, statistical methods for high-throughput biological experimentation and the dynamic modelling of cellular systems. This work is highly interdisciplinary and usually involves close collaboration with experimental biologists and clinicians. A recurrently theme is the study of complex cellular networks at ...
Projects: BaCell-SysMO
Institutions: University of Marburg
Projects: TRALAMINOL, CC-TOP
Institutions: BASF
Projects: Sustainable co-production
Institutions: University of Rostock
Projects: Not specified
Institutions: Not specified
Matexcel provides HAp in different dimensions, from nano to micro sizes, as well as different grades such as research, medical and food grades.
Projects: Xenophiles Systems Biology
Institutions: VU University Amsterdam
Professor at VU
Projects: STREAM, SCaRAB, WineSys, INBioPharm
Institutions: SINTEF, Norwegian University of Science and Technology
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: COVID-19 Disease Map
Institutions: Radboud University Medical Center
 https://orcid.org/0000-0002-5353-7691
https://orcid.org/0000-0002-5353-7691
Projects: Quantitative EEG normal values
Institutions: University Hospital Essen
Lutz Brusch is heading the research group "Spatio-temporal pattern formation in cells and tissues" at the Centre for Information Services and High Performance Computing of TU Dresden, Germany. The group is co-developing the multi-cellular modelling and simulation framework Morpheus (https://morpheus.gitlab.io) and is collaborating with experimental labs on questions of tissue morphogenesis and regeneration.
Projects: PSYSMO
Institutions: University of Dortmund
Projects: TestingSeek, Project Test, FAIRDOM
Institutions: University of Edinburgh, Institute of Physiology Academy of Sciences
Projects: SysMetEx, Kinetics on the move - Workshop 2016
Institutions: Università della Svizzera Italiana
Projects: PhotoBoost
Institutions: Fraunhofer IME
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
Projects: Genetic diversity and forage yield of perennial ryegrass, FAIRDOM Community Workers
Institutions: University of Ghent
Projects: STREAM
Institutions: University of Warwick
Projects: COVID-19 Disease Map
Institutions: Hasso-Plattner Institute/ TU Berlin
 https://orcid.org/0000-0001-8231-0501
https://orcid.org/0000-0001-8231-0501
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: PhotoBoost
Institutions: Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Projects: CC-TOP
Institutions: University of Zagreb
Projects: COVID-19 Disease Map
Institutions: Institut Curie
Projects: CC-TOP
Institutions: European Research Executive Agency
PO
Projects: HUMET Startup
Institutions: Université catholique de Louvain
 https://orcid.org/0000-0003-2040-2448
https://orcid.org/0000-0003-2040-2448
Professor Patrice D. Cani is researcher from the Belgian Fund for Scientific Research and group leader in the Metabolism and Nutrition lab at the Louvain Drug Research Institute from the UCL, Brussels, Belgium. He is WELBIO investigator and recipient of an ERC Starting Grant 2013 and a PoC ERC Grant 2016. He is laureate of the Baillet-Latour grant for medical research and the international prize of Physiology Lucien Dautrebande. His main research interests are the investigation of interactions ...
Projects: WURSynBio, INDIE - Biotechnological production of sustainable indole
Institutions: Wageningen University & Research
Projects: PhotoBoost
Institutions: University of Leida
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: Universitat Rovira i Virgili
Projects: TRALAMINOL
Institutions: University College London (UCL)
Elizabete is an expert on the regulation of carbon assimilation by Rubisco in crop plants, especially wheat and cowpea. She leads a research team that aims to understand and improve the efficiency of photosynthesis to optimise the sustainability and climate resilience of crop production.
She received her undergraduate degree in applied plant biology at the University of Lisbon, where she went on to earn her PhD researching photosynthesis and photorespiration in C4 grasses. She specialized on the ...
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: Thetis
Projects: Simulation Foundries
Institutions: University of Stuttgart
 https://orcid.org/0000-0001-8809-1856
https://orcid.org/0000-0001-8809-1856
Projects: PhotoBoost
Institutions: Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Projects: COVID-19 Disease Map
Institutions: Swiss Institute of Bioinformatics
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: Thetis
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
 https://orcid.org/0000-0002-7350-9161
https://orcid.org/0000-0002-7350-9161
Projects: NIRBTEST
Institutions: Institut Curie
Projects: COVID-19 Disease Map
Institutions: University of Rome Tor Vergata
Projects: COVID-19 Disease Map
Institutions: Policlinico Universitario Agostino Gemelli IRCCS
ESR14
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur de Tunis
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: STREAM
Institutions: University of Warwick
Projects: SulfoSys - Biotec
Institutions: University of Braunschweig
Projects: TRALAMINOL, CC-TOP
Institutions: PROZOMIX
Projects: COVID-19 Disease Map
Institutions: Shri Govindram Seksaria Institute of Technology and Science
Projects: COVID-19 Disease Map
Institutions: Centre national de la Recherche Scientifique (CNRS)
Projects: datamgmt
Institutions: insilicotrials Technologies
Projects: the Supplementary materials for paper
Institutions: The First Affiliated Hospital of Guangzhou Medical University
Projects: COVID-19 Disease Map
Institutions: Monash University
 https://orcid.org/0000-0002-9207-0385
https://orcid.org/0000-0002-9207-0385
Projects: Consensus Hallmark Annotation, FAIR Functional Enrichment, The evolution of Gene Ontology
Institutions: University of Leiden, LIACS
Projects: Millar group, TiMet
Institutions: University of Edinburgh
 https://orcid.org/0000-0002-9258-583X
https://orcid.org/0000-0002-9258-583X
Projects: Genomic Medicine
Institutions: Kyungpook National University
Projects: Molecular Systems Biology
Institutions: Stellenbosch University
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0001-6229-2211
https://orcid.org/0000-0001-6229-2211
Projects: PhotoBoost
Institutions: University of Leida
Projects: Mapping Regional Susceptibility to Neurodegenerative Diseases
Institutions: Technical University of Munich
Projects: Sustainable co-production
Institutions: Wageningen University & Research
Projects: RootBook
Institutions: ETH Zurich
Projects: TRALAMINOL, CC-TOP
Institutions: CSIC Barcelona - Consejo Superior de Investigaciones Científicas
Projects: MESI-STRAT
Institutions: University of Newcastle
Projects: SilicoTryp
Institutions: University of Heidelberg
Projects: SysMO-LAB
Institutions: Heidelberg Institute for Theoretical Studies
- BSc in Physics and Chemistry at the West University Timisoara 1999 - graduated with a average grade of 9.57 (on a scale from 1.00 to 10.00)
- Accepted in the International MSc/PhD Program Molecular Biology - International Max Planck Research School, Goettingen, Germany in June 2000. For details about the program please consult http://www.gpmolbio.uni-goettingen.de/
- Master in Molecular Biology at the Georg-August Univ. Goettingen, Germany, August 2001 - graduated with a grade of 2.0 (on a scale ...
Projects: SAFE-Aqua, Biomics Projects
Institutions: Institut Pasteur
 https://orcid.org/0000-0001-6286-1138
https://orcid.org/0000-0001-6286-1138
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: University of Milano-Bicocca
Projects: Biospecimen Collection Protocol
Institutions: creative bioarray
Creative Bioarray is an innovative biotechnology company whose mission focuses on developing unique technologies that provide global scientists with high quality products and satisfactory services to facilitate the investigation of life science researches.
Projects: COVID-19 Disease Map
Institutions: University of Florida
Projects: CC-TOP
Institutions: Technische Universität Darmstadt
Projects: Principais cânceres que afetam mulheres no Brasil
Institutions: Instituto Nacional do Câncer
Projects: COVID-19 Disease Map
Institutions: University of Bonn
Projects: COVID-19 Disease Map
Institutions: University Maastricht
Projects: OXYMOD
Institutions: Universidade Nova de Lisboa, FCT-NOVA
 https://orcid.org/0000-0002-7892-8955
https://orcid.org/0000-0002-7892-8955
Projects: SCaRAB
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: NMTrypI - New Medicines for Trypanosomatidic Infections, Mass spectrometry proteomics for biomarker discovery, Thymidylate synthase dimer dissociation, BioTarget Database, Compound Data, Giulia Malpezzi PhD raw data, Defining the role of fibroblasts in skin expansion, Thymidylate synthase dimer disrupters induce DNA damage, halt cell growth and overcome drug resistance in colorectal cancer
Institutions: Università degli Studi di Modena e Reggio Emilia (UNIMORE), Life Science, University of Modena and Reggio Emilia
 https://orcid.org/0000-0002-0443-5402
https://orcid.org/0000-0002-0443-5402
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-1644-3223
https://orcid.org/0000-0002-1644-3223
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur
Projects: Dynamic interplay between nuclei and mitochondria in ageing cells
Institutions: National Institute on Aging
Projects: SUMO
Institutions: University of Edinburgh
Projects: COSMIC
Institutions: Beuth University of Applied Sciences Berlin
I am a biotechnologist with main focus on theoretical studies. Currently, I am working on the implementation of a parameter estimation algorithm on GPUs to reduce the computational burden of huge ODE systems. I am a PAL and I am looking forward to communication with other SYSMO members.
Projects: TRANSLUCENT
Institutions: University of Cordoba
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Gothenburg
Projects: Not specified
Institutions: Not specified
Alfa Cytology is a premier firm specializing in cancer research and therapeutic development, armed with state-of-the-art technology and a team of renowned scientists. The company offers tailored services in tumor research, committed to delivering high-quality, innovative, and efficient solutions. Alfa Cytology has been striving to expand and improve itself, always providing customers with reliable and scientific services.
Projects: COVID-19 Disease Map
Institutions: University of Applied Sciences Mittweida
 https://orcid.org/0000-0002-1788-9593
https://orcid.org/0000-0002-1788-9593
Projects: COVID-19 Disease Map
Institutions: Yenepoya University
Projects: COVID-19 Disease Map
Institutions: NYU Grossman School of Medicine
 https://orcid.org/0000-0002-5494-626X
https://orcid.org/0000-0002-5494-626X
Projects: SysVirDrug
Institutions: German Cancer Research Center (DKFZ)
Projects: SteaPKMod, Surgery Analyzer, Quantifizierung des Zusammenhanges zwischen Leberperfusion und -funktion bei erweiterter Leberresektion - Ein systemmedizinischer Ansatz (QuaLiPerF)
Institutions: Experimental Transplantation Surgery, Department of General, Visceral and Vascular Surgery, University Hospital Jena, Experimental Transplantation Surgery, Jena University Hospital
 https://orcid.org/0000-0003-3483-3388
https://orcid.org/0000-0003-3483-3388
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: SCaRAB
Institutions: University of Greifswald
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Projects: SysMO-LAB
Institutions: University of Heidelberg
I am working on the development of algorithms for Comparative Systems Biology.
Institutions: CropXR, Utrecht University
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0002-8839-3333
https://orcid.org/0000-0002-8839-3333
Projects: Winter Wheat (Triticum aestivum L.) Grain Yield, Quality, and Net Photosynthesis When Grown Under Semi-Transparent Cadmium Telluride Photovoltaic Modules Near Maturity, Impacts of a Vertical Bifacial Agrivoltaics System on Field Corn in Northern Colorado, USA
Institutions: Colorado State University
 https://orcid.org/0009-0002-2133-1784
https://orcid.org/0009-0002-2133-1784
Projects: Cell4Chem
Institutions: Centre national de la Recherche Scientifique (CNRS)
Projects: IMOMESIC
Institutions: Griffin Discoveries BV
Projects: Sustainable co-production
Institutions: Sunchem Italy
Projects: BaCell-SysMO
Institutions: University of Groningen
Projects: NIRBTEST
Institutions: Amsterdam UMC
Projects: WATCHER
Institutions: Universidade de Santiago de Compostela
Data Steward for the Plant Biology and Breeding Division of INRAe (https://www.inrae.fr/en/divisions/bap)
Projects: PSYSMO, EmPowerPutida, SynBio4Flav
Institutions: CSIC Madrid, CSIC
Projects: Light and plant development
Institutions: University of Edinburgh
Projects: PTPN11 mutagenesis
Institutions: Josep Carreras Leukaemia Research Institute (IJC)
 https://orcid.org/0000-0001-7325-0631
https://orcid.org/0000-0001-7325-0631
Projects: COSMIC, SysMO-LAB, HUMET Startup
Institutions: Wageningen University & Research
Anita de Waard (she/her) Vice President of Research Collaborations Elsevier Research Collaborations Unit 71 Hanley Lane, Jericho, VT 05465 @anitawaard | +1 (619) 252 8589
Projects: INBioPharm
Institutions: SINTEF
Projects: SysMO-LAB
Institutions: Technical University of Denmark - Systems Biology
Projects: Vitis Data Crop
Institutions: Institut Français de la Vigne et du Vin
 https://orcid.org/0000-0001-7278-3209
https://orcid.org/0000-0001-7278-3209
Projects: COVID-19 Disease Map
Institutions: Oregon Health & Science University (OHSU)
Projects: EmPowerPutida
Institutions: University of Stuttgart
Projects: reNEW
Institutions: BMI/reNEW, University of Copenhagen
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Leiden
Position : Professor, Centre of Integrative Genomics, Faculty of Biology and Medicine, University of Lausanne
My research interests focus on understanding metabolic homeostasis. We are mainly using genetically modified mouse models and systems approaches. I am also very keen in exploring how Omics approaches is changing, or not changing, key biological concepts. Link to complete publication list : http://orcid.org/0000-0001-5483-288X
Projects: COVID-19 Disease Map
Institutions: Lorraine Research Laboratory in Computer Science and its Applications (LORIA)
Projects: MESI-STRAT
Institutions: German Cancer Research Center (DKFZ)
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: BESTER
Institutions: Imperial College London
Projects: CC-TOP
Institutions: University of Goettingen
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: SysVirDrug
Institutions: ETH Zurich
Projects: STREAM
Institutions: University of Groningen
Projects: Hi-IMPAcTB, MIT SRP, MetNet
Institutions: Massachusetts Institute of Technology
Projects: SysVirDrug
Institutions: ETH Zurich
Projects: Bio-crop
Institutions: Paris Brain Institute / Inria
Projects: HUMET Startup, COVID-19 Disease Map
Institutions: Bilkent University
 https://orcid.org/0000-0002-7153-0784
https://orcid.org/0000-0002-7153-0784
Christin is a project manager at DOCYET. She has a degree in social work, public health and medical ethics. In addition to her full-time professional work, she is a PhD candidate at the University of Bremen. She has training and certificates in data stewardship, project & change & hospital management. The following research and application areas fascinate her mostly: health-related data/infrastructure, FAIRification, robotics in medicine, health standards and digital health solutions.
Projects: EmPowerPutida
Institutions: Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Projects: Xenophiles Systems Biology
Institutions: University of Amsterdam
Professor of Astrophysics at the University of Amsterdam
Projects: BESTER
Institutions: Imperial College London
My scientific interests revolve around functional genomics, systems biology and the development of algorithms and software for the analysis of high-throughput data (mainly, but not restricted to, Next Generation Sequencing) and its application to the relationship between genotype and phenotype, mainly oriented to personalized and precision medicine. I am especially interested in the study of disease mechanisms and drug action mechanisms, drug repositioning and the definition of mechanism-based ...
Projects: SysMetEx
Institutions: Linnaeus University
Projects: HOTSOLUTE
Institutions: Consiglio Nazionale delle Ricerche
 https://orcid.org/0000-0002-7766-2040
https://orcid.org/0000-0002-7766-2040
Projects: COSMIC
Institutions: University of Goettingen
Projects: NIRBTEST
Institutions: Amsterdam UMC
Projects: NIRBTEST
Institutions: Institut Curie
Projects: iRhythmics
Institutions: Leibniz Institute for Farm Animal Biology (FBN)
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0002-1240-5553
https://orcid.org/0000-0002-1240-5553
Andreas Dräger is the assistant professor for Computational Systems Biology of Infection and Antimicrobial-Resistant Pathogens at the University of Tübingen in Germany. His group aims to combat the spreading antibiotics resistances by using mathematical modeling and computer simulation of bacterial systems up to entire microbiomes and host-pathogen interactions. In doing so, his group actively contributes to the advancement of various COMBINE standards.
Projects: SulfoSys
Institutions: University of Groningen
Projects: COVID-19 Disease Map
Institutions: University of Lyon
Projects: SysMO DB
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: SABIO-VIS
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0001-5034-730X
https://orcid.org/0000-0001-5034-730X
Projects: COVID-19 Disease Map
Institutions: University of Heidelberg
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Buenos Aires
Projects: Siduri (test)
Institutions: INRAE
Projects: PSYSMO
Institutions: CSIC Madrid
Institutions: University of Ulm
Prof. Dr. Peter Dürre Head, Department of Microbiology and Biotechnology, University of Ulm 89069 Ulm, Germany
Assistant Professor, Department of Biochemistry, All India Institute of Medical Sciences Kalyani
Projects: COVID-19 Disease Map
Institutions: Oregon State University
Projects: Bio4Fuels
Institutions: Norwegian University of Life Sciences
Projects: COVID-19 Disease Map
Institutions: University of Pittsburgh
Projects: DigiSal-BT8121
Institutions: Inland Norway University of Applied Sciences
 https://orcid.org/0000-0003-3134-5186
https://orcid.org/0000-0003-3134-5186
Projects: SUMO
Institutions: University of Stuttgart
I'm interested in the application and development of methods of systems theory in biology (systems biology). In particulary I work on the following topics: Thermodynamic constraints on biochemical network; Model reduction; Modeling and Analysis of metabolic regulation.
Projects: CC-TOP
Institutions: University of Goettingen
Projects: MESI-STRAT
Institutions: Medical University Innsbruck
Projects: COSMIC
Institutions: University of Goettingen
I have a permanent position at the department of microbiology at the TU-München. As a microbiologist I am interested in the regulation of central metabolism in prokaryotic organisms with different types of energy metabolism such as Clostridia, Bacilli and acetic acid bacteria. Furthermore I worked as a software developer for several years in a bioinformatics company and I am very interested in bioinformatics and handling of large amounts of data.
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Humboldt-Universität zu Berlin
I am working at the boundary of wet-labs and mathematical modeling, trying to understand the regulation of the eukaryotic cell cycle. I started working with yeast in a group of Systems Biology (Prof. Edda Klipp, Humboldt University, Berlin). Before that I studied biology, engineering and did research on the nuclear pore complex.
Projects: COVID-19 Disease Map
Institutions: University Maastricht
 https://orcid.org/0000-0002-7770-620X
https://orcid.org/0000-0002-7770-620X
Projects: Systems toxicology of Atlantic cod, DigiSal
Institutions: University of Bergen
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-9220-8743
https://orcid.org/0000-0002-9220-8743
Projects: COVID-19 Disease Map
Institutions: University Maastricht
Projects: SysMetEx
Institutions: University Duisburg-Essen
Projects: MESI-STRAT
Institutions: PD-Value
Jeroen Elassaiss-Schaap, (jeroen@pd-value.com)
Projects: COVID-19 Disease Map
Institutions: University of Lille
 https://orcid.org/0000-0002-4440-2904
https://orcid.org/0000-0002-4440-2904
Projects: sample project
Institutions: de
Projects: SulfoSys
Institutions: University of Bergen
Projects: AquaHealth (ERA-BlueBio)
Institutions: Sea & Sun Technology GmbH
Projects: STREAM, SCaRAB, INBioPharm, SYSTERACT
Institutions: SINTEF
Projects: MRC-UNICORN
Institutions: Imperial College London
Projects: MESI-STRAT, MESI-STRAT Review, MESI-Review 2024
Institutions: University of Innsbruck
Projects: 3T3 murine fibroblast as feeder-layer, Biochemical characterization of the feedforward loop between CDK1 and FOXM1 in epidermal stem cells, PRIN 2022 - High-resolution study of translational dynamics in human epithelial stem cells, Defining the role of fibroblasts in skin expansion
Institutions: Università degli Studi di Modena e Reggio Emilia (UNIMORE), Università di Modena e Reggio Emilia, Life Science, University of Modena and Reggio Emilia
 https://orcid.org/0000-0001-9768-6368
https://orcid.org/0000-0001-9768-6368
Projects: WG Infrastructure for Translational Research, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", Translational Bioinformatics, Medical Biometry, Epidemiology, Medical Informatics
Institutions: University Medical Center Göttingen, University of Tübingen, University of Saarland
 https://orcid.org/0000-0002-1505-594X
https://orcid.org/0000-0002-1505-594X
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
I'm currently a Postdoc at the Institute of Technical Biochemistry in Stuttgart University. My project involves the experimental validation of the Indirect Enzymatic Dehydration Via Phosphorylation and Dephosphorylation of Isobutanol for Isobutene production.
Projects: COVID-19 Disease Map
Institutions: Fundación Progreso y Salud
 https://orcid.org/0000-0003-2632-9587
https://orcid.org/0000-0003-2632-9587
Projects: HubMOL
Institutions: University Hospital Essen
Projects: Viral Metagenomic
Institutions: CIRAD
Projects: SilicoTryp
Institutions: University of Heidelberg
Projects: COVID-19 Disease Map
Institutions: University of Lisbon
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: John Innes Centre
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: Thetis
F.E. Faradjev is a prominent Soviet, Azerbaijani and Swedish scientist and educator. He is one of the pioneers of nanophotonics. In 2025, he was elected as a Corresponding Member of the Russian Academy of Natural Sciences and has recently been awarded the honorary title "Honored scientist and educator" for developing priority scientific areas, establishing schools, and his contributions to STEM education.
Projects: FUNCEMM
Institutions: BOKU University
Projects: C19DM - Macrophage logical model, COVID-19 Disease Map
Institutions: Norwegian University of Science and Technology
Projects: COVID-19 Disease Map
Institutions: Forman Christian College (a chartered university)
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: DigiSal
Institutions: Norwegian University of Life Sciences
Projects: SysMO-LAB
Institutions: Heidelberg Institute for Theoretical Studies, University of Heidelberg
Projects: ROBUSTYEAST
Institutions: Ecole Polytechnique Fédérale de Lausanne (EPFL)
 https://orcid.org/0000-0001-8110-8424
https://orcid.org/0000-0001-8110-8424
Projects: COVID-19 Disease Map
Institutions: University of Edinburgh
Projects: C1Pro
Institutions: Norwegian University of Science and Technology
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: MPW-Val
Institutions: Universidade de Santiago de Compostela
Projects: HOTSOLUTE
Institutions: Consiglio Nazionale delle Ricerche
 https://orcid.org/0000-0002-3390-9638
https://orcid.org/0000-0002-3390-9638
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Bergen
Projects: Toggle switch, Reduce Complexity (RCO) reconstruction
Institutions: Heinrich Heine University of Düsseldorf
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: TRALAMINOL, CC-TOP
Institutions: Technische Universität Darmstadt
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: SysMO-LAB
Institutions: University of Rostock
Postdoc @ Institute of Medical Microbiology, University of Rostock
Projects: SulfoSys - Biotec
Institutions: Otto-von-Guericke-Universität Magdeburg
Projects: Thesis student-FrancescaFilippelli
Institutions: Università degli Studi di Modena e Reggio Emilia (UNIMORE)
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: CC-TOP
Institutions: University of Zagreb
Projects: EmPowerPutida
Institutions: University of Stuttgart
Projects: INCOME
Institutions: ICB Helmholtz Center Munich
Projects: COSMIC
Institutions: University of Rostock
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Leiden
Projects: Millar group
Institutions: University of Edinburgh
Projects: CausalDB, Colosys, NTNU Health Druglogics
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-3357-425X
https://orcid.org/0000-0002-3357-425X
Projects: BaCell-SysMO
Institutions: University of Goettingen
I'm a PhD student at the lab of Prof. Dr. Jörg Stülke. My main interest is to analyze the central metabolism of Bacillus subtilis using systems biology software. I have developed an algorithm to find short pathways connecting sets of metabolites and I'm also involved in SubtiWiki, the wiki for all genes of Bacillus subtilis (http://subtiwiki.uni-goettingen.de)
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München / Institute of Experimental Genetics
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Duesseldorf
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-8236-7986
https://orcid.org/0000-0002-8236-7986
Projects: MPIEvolBio-SciComp
Institutions: Max Planck Institute for Evolutionary Biology
 https://orcid.org/0000-0002-2579-5546
https://orcid.org/0000-0002-2579-5546
I'm heading the Scientific Computing Unit at Max Planck Institute for Evolutionary Biology in Plön, Germany. My interest is the application of high performance and high throughput computing in data processing, data analysis, and numerical simulations of dynamical processes. I have a background in Theoretical Physics with a Ph.D. from Rostock University (2008), where I studied analytical models and numerical simulations of extremely dense and hot matter.
After a postdoc at the Lawrence Livermore ...
Projects: EmPowerPutida
Institutions: Ingenza Ltd.
Projects: HUMET Startup
Institutions: Universitair ziekenhuis Antwerpen
 https://orcid.org/0000-0002-7527-4714
https://orcid.org/0000-0002-7527-4714
Sven Francque is currently Head of the Department of Gastroenterology and Hepatology of the Antwerp University Hospital and full professor of gastroenterology and hepatology at the University of Antwerpen, with a longtanding interest and expertise in the field of Non-Alcohloic Fatty Liver Disease. In collaboration with the Department of Endocrinology, Diabetes and Metabolic Diseases, a large cohort of well-phenotyped patients has been built up.
Projects: PSYSMO
Institutions: University of Hannover
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: Roche Diagnostics
Projects: COVID-19 Disease Map
Institutions: University of Edinburgh
Projects: COVID-19 Disease Map
Institutions: University of Edinburgh
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0001-8003-6860
https://orcid.org/0000-0001-8003-6860
Research Scientist, Computer Vision; Intellisense Systems, Inc. Lead Developer for MultiCellDS
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München / Institute of Experimental Genetics
Projects: TRANSLUCENT
Institutions: University of Vienna
Projects: PSYSMO
Institutions: University of Duesseldorf
Projects: COVID-19 Disease Map
Institutions: Hospital del Mar Research Institute (IMIM)
 https://orcid.org/0000-0002-9383-528X
https://orcid.org/0000-0002-9383-528X
Projects: COVID-19 Disease Map
Institutions: Pondicherry University
 https://orcid.org/0000-0003-4854-8238
https://orcid.org/0000-0003-4854-8238
Assistant Professor (Research) at Department of Bioinformatics, SASTRA University.
Projects: KOSMOBAC
Institutions: Ludwig-Maximilians University of Munich
I am a PhD student of the microbiology department at the Ludwig-Maximilians Universität München. I work at the chair of Prof. Kirsten Jung. The topic of our workpackage deals with "K+ homeostasis in Escherichia coli". In special I'm working on the sensor kinase KdpD that controls together with the response regulator KdpE the expression of the high-affinity K+ uptake system KdpFABC. The yet not fully understood molecular mechanism of stimulus perception and signal transduction is of particular ...
Projects: MESI-STRAT
Institutions: University of Heidelberg
Projects: FAIRDOM user meeting
Institutions: UPF
Projects: Enzymatic biorecycling of PET (PETzyme)
Institutions: Universidade de Santiago de Compostela
Projects: iRhythmics
Institutions: Leibniz Institute for Farm Animal Biology (FBN)
 https://orcid.org/0000-0002-7968-3152
https://orcid.org/0000-0002-7968-3152
Projects: CF transcriptome, miRiAD - exploring the role of microRNAs in T cell function and anti-viral defence, ComPASs - Common Pathways in Amyotrophic Lateral Sclerosis (ALS) and Spinal Muscular Atrophy (SMA), LungCARD - Blood test for clinical therapy guidance of non-small cell lung cancer patients, UnCentre - Unlocking satellite DNA and centromere structure in vertebrate chromosomes, PhotoBoost
Institutions: RNA Systems Biology Lab - BioISI/FCUL, Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Group Leader and Assistant Professor
Projects: Noisy-Strep
Institutions: University of Groningen
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Institutions: Universidade de Santiago de Compostela
Projects: LEANPROT
Institutions: Norwegian University of Science and Technology
Projects: KOSMOBAC
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: STREAM
Institutions: University of Warwick
Projects: INCOME
Institutions: University of Rostock
Projects: SynBio4Flav
Institutions: Symrise
Projects: COVID-19 Disease Map
Institutions: Mediterranean Institute for Life Sciences (MedILS)
Projects: Not specified
Institutions: Not specified
CD Genomics is a leading global life sciences company, and we remain committed to providing the research community with high-quality long-read sequencing services, from Oxford Nanopore to PacBio SMRT sequencing.
Projects: WG1 - Compound libraries coordination and integration of compound design
Institutions: University of Siena
Projects: TRANSLUCENT
Institutions: Humboldt-Universität zu Berlin
Projects: SulfoSys
Institutions: University of Braunschweig
Projects: SCaRAB
Institutions: University of Greifswald
Projects: Lipopeptide research @ VIB
Institutions: VIB
Projects: COVID-19 Disease Map
Institutions: Moredun Research Institute
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur de Tunis
Yousof Gheisari is focused on developing novel therapeutics for chronic kidney disease, with a particular emphasis on diabetic kidney disease. To achieve this goal, his research group uses systems biology and artificial intelligence approaches for multi-omics analysis, big-data integration, large-scale dynamic modeling, drug-target prioritization, in-silico lead generation, and drug repurposing. Predictions from these pipelines are subsequently assessed in preclinical models. The interdisciplinary ...
Projects: Japanese Data in Global Plant Research: A FAIDARE and BrAPI Project
Institutions: National Institute of Genetics
Institutions: University of Gothenburg, European Molecular Biology Laboratory
Projects: CoolWine
Institutions: University of Gothenburg, European Molecular Biology Laboratory
Projects: TERRACOTA
Institutions: Systems Biomedicine Center, University of Luxembourg
Research Associate, SDBV
Projects: S. cerevisiae fed-batch fermentation
Institutions: RWTH Aachen University
Projects: PROJECT___DR_Levels1
Institutions: DBMI
Projects: Master-BIDS, PROJECT___DR_Levels1, PROJECT__DR_Level2, DR July
Institutions: DBMI, University Medical Centre Mannheim
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: EnzymeML
Institutions: Technische Universität Berlin (TU Berlin)
 https://orcid.org/0000-0002-0254-1500
https://orcid.org/0000-0002-0254-1500
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-9533-3227
https://orcid.org/0000-0001-9533-3227
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
 https://orcid.org/0000-0002-5766-1702
https://orcid.org/0000-0002-5766-1702
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: Remodeling of cIV
Institutions: University of Giessen
 https://orcid.org/0000-0001-6315-7361
https://orcid.org/0000-0001-6315-7361
I am performing research in biomarkers discovery, particularly regarding liver fibrosis in non-alcoholic fatty liver disease. My research activities involved the use of bioinformatic platforms, the analysis of biological protein-protein interaction networks and the experimental validation of candidates by ELISA.
Projects: OXYMOD
Institutions: National Laboratory of Energy and Geology (LNEG)
 https://orcid.org/0000-0002-5376-8430
https://orcid.org/0000-0002-5376-8430
Projects: SilicoTryp
Institutions: University of Glasgow
Projects: TRALAMINOL
Institutions: PROZOMIX
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
Projects: MOSES
Institutions: Medical University Vienna
General Bioinformatics
Projects: Subject collection
Institutions: University of Freiburg
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0003-3634-190X
https://orcid.org/0000-0003-3634-190X
Dr. Glazier’s research focuses on early embryonic development, developmental and chronic toxicity and disease, with more than 100 experimental and computational papers on biological development and developmental diseases (including polycystic kidney disease (ADPKD), tumor growth and vascularization, Age Related Macular Degeneration and diabetic retinopathies, somitogenesis and liver toxicity) and more recently on modeling in-host viral infection and immune response. As part of his work on infection ...
Projects: COMBINE Multicellular Modelling
Institutions: University College London (UCL)
 https://orcid.org/0000-0001-5963-8576
https://orcid.org/0000-0001-5963-8576
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Professor of Computer Science University of Manchester Co-Director of the FAIRDOM Initiative and co-leader of the SEEK4Science Platform Development Deputy Head of Node ELIXIR-UK Co-lead ELIXIR Interoperability Backbone Platform Lead ISBE WP Data and Model Management Data lead SynBioChem Manchester Synthetic Biology Research Centre for Fine and Speciality Chemicals
Projects: HubMOL
Institutions: Consiglio Nazionale delle Ricerche
 https://orcid.org/0009-0006-0802-5853
https://orcid.org/0009-0006-0802-5853
Projects: Data science Übung
Institutions: Technical University of Munich
Projects: MESI-STRAT
Institutions: University of Bergen
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: Iceni Diagnostics Ltd
Projects: COVID-19 Disease Map
Institutions: KAUST
Projects: COVID-19 Disease Map
Institutions: University of Western Australia
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0003-0073-3282
https://orcid.org/0000-0003-0073-3282
Projects: COVID-19 Disease Map
Institutions: ICAHN Icahn School of Medicine at Mount Sinai
Projects: FAIRDOM, Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, MS_DILI, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training, EnzymeML, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", FAIRDOM Community Workers, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, ModeleXchange initiative, SDBV/HITS, EDITH (Ecosystem Digital Twins in Health) test project
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-8683-7084
https://orcid.org/0000-0002-8683-7084
Data management and standardization expert for systems biology and systems medicine, responsible for the data management user requirements and user contacts within the German LiSyM network (Liver Systems Medicine: http://lisym.org/) and associated to the FAIRDOM team. Involved in different standardization initiatives and committees, i.e. COMBINE (http://co.mbine.org), ISO/TC 276 Biotechnology (https://www.iso.org/committee/4514241.html), European COST action CHARME (http://www.cost-charme.eu) and ...
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Projects: CropClock
Institutions: Systems Biomedicine Center, University of Luxembourg
 https://orcid.org/0000-0002-5228-6165
https://orcid.org/0000-0002-5228-6165
Projects: COVID-19 Disease Map
Institutions: Institute of Bioinfomratics
Projects: KOSMOBAC
Institutions: Max-Planck-Institute of Biochemistry Martinsried
Professor in Jinan University, Guangzhou, China. My research interest is in the modeling of translation. Connecting various processes in translation, we can investigate the impact of different factors on protein biosynthesis and biogenesis in genome-wide scale. This may reveal various general mechanisms on control level of gene expression and folding efficiency regulation in different growth conditions.
He earned his degree in Pharmacy in 2009 from the University of Granada, receiving a national collaboration grant in his fifth year to work in the Department of Microbiology. Later, through an excellence contract from the Andalusian Regional Government, he completed his international doctoral thesis at the University of Granada, studying new autotrophic nitrogen removal systems from both a technical and microbiological perspective. As part of his doctoral research, he conducted a research stay ...
Projects: SCaRAB
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: SynBio4Flav
Institutions: German Institute of Human Nutrition
 https://orcid.org/0000-0002-9977-5994
https://orcid.org/0000-0002-9977-5994
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: XyloCut
Institutions: Lund University
Projects: FAIRDOM user meeting
Institutions: Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences
Projects: COSMIC
Institutions: Beuth University of Applied Sciences Berlin
 https://orcid.org/0000-0001-6096-1354
https://orcid.org/0000-0001-6096-1354
Process engineer, modeling biological systems since 1985.
Projects: Kinetics on the move - Workshop 2016
Institutions: Novosibirsk State University
 https://orcid.org/0000-0001-8957-6142
https://orcid.org/0000-0001-8957-6142
Projects: SUMO
Institutions: University of Sheffield
Post-doctoral research associate working in Sheffield in the SUMO consortium.
Projects: TRALAMINOL
Institutions: PROZOMIX
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-9551-9280
https://orcid.org/0000-0002-9551-9280
Projects: TRANSLUCENT
Institutions: University of Vienna
Projects: Not specified
Institutions: Not specified
CD Bioparticles is a leading manufacturer and supplier of various nanoparticles, microparticles, and coatings for R&D as well as commercialization across different application areas, including in vitro diagnostics, biochemistry, cellular analysis, cell separation, and immunoassay. The company also offers various custom services, including chemical surface-functionalization, fluorescent modification, antibody immobilization, as well as nucleic acid and oligo ...
Projects: Not specified
Institutions: Not specified
Hi, this is Caroline from Creative Biomart, a world leading biotech company that specialized in providing protein, recombinant protein, cell lines, and GMP proteins. visit http://www.creativebiomart.net/ to see more details.
Projects: Not specified
Institutions: Not specified
Creative BioMart offers comprehensive, one-stop services across the entire protein production workflow including gene cloning and synthesis to protein analytical.
Projects: SUMO
Institutions: University of Sheffield
The major theme of the research in my laboratory is bacterial gene regulation. We are interested in signal perception mechanisms (in particular oxygen); signal transduction (ligand induced protein confromational changes); interaction of transcription factors with the core transcription machinery; interactions between transcription factors to integrate multiple signals; and the influence of promoter architectures on these events. We are also interested in aome aspects of post-transcriptional ...
Projects: KOSMOBAC
Institutions: University of Aberdeen
Projects: Stress granules, MESI-STRAT
Institutions: University of Bergen
More detailed profile here https://www.uib.no/en/persons/Sushma.Grellscheid
Projects: Rhodolive
Institutions: National Institute of Chemistry (Slovenia)
 https://orcid.org/0000-0002-8255-647X
https://orcid.org/0000-0002-8255-647X
Projects: COVID-19 Disease Map
Institutions: Biomathematica
POSITION Professor of Systems Medicine/Biology at the Academic Medical Center, Amsterdam and University Medical Center Groningen, Groningen, The Netherlands
RESEARCH My major projects focus on understanding the etiology of metabolic syndrome and its comorbidities type2 diabetes and cardiovascular disease. To get grip on the sequence of events in disease progression we make use of longitidunal models and apply multiscal systems biology approaches.
PUBLICATIONS
My published work can be found at: ...
Projects: Not specified
Institutions: Not specified
Professor of Artificial Intelligence at Tehnical University of Cluj-Napoca, Romania https://scholar.google.com/citations?user=xnB-QzIAAAAJ&hl=en https://users.utcluj.ro/~agroza/
Projects: HYp - Spatiotemporal analysis of hypersensitive response to Potato virus Y in potato, pISA-tree, MOA - Multiomics analysis of potato response to Potato virus Y (PVY) infection, SUSPHIRE - Sustainable Bioproduction of Pheromones for Insect Pest Control in Agriculture, INDIE - Biotechnological production of sustainable indole, FAIRDOM user meeting, _p_stRT, ADAPT - Accelerated Development of multiple-stress tolerAnt PoTato
Institutions: National Institute of Biology
 https://orcid.org/0000-0001-5906-8569
https://orcid.org/0000-0001-5906-8569
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: Lobet's group
Institutions: Forschungszentrum Jülich
 https://orcid.org/0000-0002-5883-4572
https://orcid.org/0000-0002-5883-4572
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: DigiSal, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-5680-4855
https://orcid.org/0000-0002-5680-4855
Projects: COVID-19 Disease Map
Institutions: Imperial College London
Projects: COVID-19 Disease Map
Institutions: University of Rostock
 https://orcid.org/0000-0002-3470-3260
https://orcid.org/0000-0002-3470-3260
Projects: COVID-19 Disease Map
Institutions: University of Würzburg
Projects: KOSMOBAC
Institutions: University of Aberdeen
Polyglot European Scientist. I thrive working in interdisciplinary environments combining the study of enzyme reactions and mechanisms with bioinformatics, molecular modelling, automated data analysis and data stewardship.
Projects: COVID-19 Disease Map
Institutions: Harvard Medical School
Projects: SilicoTryp, IMOMESIC
Institutions: University of Groningen, VU University Amsterdam
Projects: PSYSMO
Institutions: University of Braunschweig
Projects: CropClock
Institutions: University of Cyprus
Projects: ExtremoPharm
Institutions: Institut National de la Recherche Agronomique
Projects: SulfoSys
Institutions: University Duisburg-Essen
Projects: AquaHealth (ERA-BlueBio)
Institutions: SINTEF
Projects: SysMetEx
Institutions: Linnaeus University
Projects: SYLOBIO
Institutions: University Medical Center Rostock, Institute of Cell Biology
 https://orcid.org/0009-0009-8356-4402
https://orcid.org/0009-0009-8356-4402
Projects: Uni Heidelberg URZ-Demo Project
Institutions: University of Heidelberg
 https://orcid.org/0009-0006-8685-3507
https://orcid.org/0009-0006-8685-3507
Projects: PSYSMO
Institutions: Queen's University Belfast
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur de Tunis
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: BaCell-SysMO
Institutions: University of Newcastle
Projects: Sustainable co-production
Institutions: Sunchem Italy
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Leiden
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: LEANPROT
Institutions: Technische Universität Berlin (TU Berlin)
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-5322-0192
https://orcid.org/0000-0002-5322-0192
Projects: COVID-19 Disease Map
Institutions: Gladstone Institutes
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Universität Konstanz
I studied Life Science (which is similar to chemical biology) at the University of Konstanz and became interested in Bioinformatics, Systems Biology and quantitative analyses during my Master's. In my PhD project I combine experimental analyses with modeling and parameter estimation approaches to quantitatively analyse regulation of apoptosis at the level of the Bcl-2 protein family.
Projects: BESTER
Institutions: Green Biologics Ltd
Projects: EmPowerPutida
Institutions: LifeGlimmer GmbH
Computational Biologist and App Designer @LifeGlimmer
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Leiden
Projects: COVID-19 Disease Map
Institutions: Institute for Globally Distributed Open Research and Education - IGDORE
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-4882-2188
https://orcid.org/0000-0003-4882-2188
Projects: BaCell-SysMO
Institutions: University of Newcastle
Optimisation of Bacillus subtilis for the secretion of heterologous proteins Therapeutic proteins (including those required for experimental purposes and clinical trials) are major products of biomanufacturing processes and considerable time and expense are expended to maximise the yield and quality of proteins produced in heterologous hosts. The production host of choice is the Gram-negative bacterium Escherichia coli for which many strains and expression systems have been developed. However, ...
Projects: INCOME, COVID-19 Disease Map
Institutions: ICB Helmholtz Center Munich, University of Bonn
 https://orcid.org/0000-0002-4935-3312
https://orcid.org/0000-0002-4935-3312
I studied Engineering Cybernetics at the University of Stuttgart and the University of Wisconsin, Madison. After my graduation, I started my PhD studies in systems biology for which I received a Ph.D. degree in 2013. A few months later I became team leader at the Institute of Computational Biology at the Helmholtz Zentrum München. Since August 2015, I lead an independent junior research group at the Helmholtz Zentrum München.
My research focuses on the development of methods for the data-driven ...
Projects: COVID-19 Disease Map
Institutions: University of Bonn
Projects: TRANSLUCENT
Institutions: University of Bonn
I am the data and model manager for the Digital Salmon
Projects: ROBUSTYEAST
Institutions: Ecole Polytechnique Fédérale de Lausanne (EPFL)
Projects: EmPowerPutida
Institutions: University of Stuttgart
Projects: Sustainable co-production
Institutions: Leuphana University of Lüneburg
 https://orcid.org/0000-0002-6112-8898
https://orcid.org/0000-0002-6112-8898
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6562-5317
https://orcid.org/0000-0001-6562-5317
Projects: C1Pro
Institutions: UNIBI: Bielefeld University
Projects: COSMIC
Institutions: University of Rostock
biomathematician, PhD student at the University of Rostock, Systems Biology Group Rostock
Projects: IDIM
Institutions: University of Freiburg
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian Institute for Water Research
 https://orcid.org/0000-0002-2981-5452
https://orcid.org/0000-0002-2981-5452
Projects: PlaSMo model repository
Institutions: University of Edinburgh
Projects: BaCell-SysMO
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: PhotoBoost
Institutions: University of Leida
Projects: COSMIC
Institutions: University of Nottingham
I'm an 'experimentalist' (molecular microbiologist) Postdoc working on regulation and peptide signaling in Clostridium acetobutylicum. I'm also a SysMO-DB PAL (Product Application Liason) for COSMIC, working on data management including standards and integration with SysMO SEEK.
Projects: BaCell-SysMO
Institutions: University of Greifswald
Projects: TRALAMINOL, CC-TOP
Institutions: Université Clermont Auvergne (UCA)
Projects: HUMET Startup
Institutions: University Medical Center Hamburg - Eppendorf Hamburg
Expert in lipid metabolism and adipose tissue
Projects: KOSMOBAC
Institutions: Ludwig-Maximilians University of Munich
I am research assistant in the microbiology department at the Ludwig-Maximilians Universität in Munich (München), working at the chair of Prof. Kirsten Jung. In our SysMO consortium we generate biological data and work in close cooperation with the workgroup of Dr. Andreas Kremling of the Max-Planck-Institut für Dynamik komplexer technischer Systeme in Magdeburg who performs mathematical modeling. The topic of our workpackage deals with "K+ homeostasis in Escherichia coli", wherby the K+ transporters, ...
Projects: MetApp, INBioPharm, SYSTERACT, Auromega
Institutions: SINTEF
 https://orcid.org/0000-0002-1221-8598
https://orcid.org/0000-0002-1221-8598
Research scientist
Projects: FAIRDOM user meeting
Institutions: University of Edinburgh
Projects: MESI-STRAT, NAD COMPARTMENTATION, NAD phylogeny, HubMOL
Institutions: The Arctic University of Norway (UiT)
 https://orcid.org/0000-0002-9124-5112
https://orcid.org/0000-0002-9124-5112
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0002-7440-2905
https://orcid.org/0000-0002-7440-2905
Research Associate in the Macklin Lab, School of Informatics, Computing, and Engineering. Indiana University, Bloomington, IN USA.
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
 https://orcid.org/0009-0008-9902-813X
https://orcid.org/0009-0008-9902-813X
Projects: BioCreative VII
Institutions: Heidelberg Institute for Theoretical Studies
Projects: DeepHyperSpec
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6726-1692
https://orcid.org/0000-0001-6726-1692
Projects: COVID-19 Disease Map, Covid-19 Interferon pathway modelling and analysis
Institutions: University of Nebraska-Lincoln
Projects: SUMO
Institutions: University of Amsterdam
General microbiologist with specific interests in signal transduction mechanisms, biophysics and photobiology
Projects: TRANSLUCENT
Institutions: University of Leiden
Projects: SUMO
Institutions: University of Stuttgart
Former: PhD student as research associate at the Institute for System Dynamics (ISYS), Universität Stuttgart, Germany. Engineering background→modelling, identification and analyses. Detailed kinetic modelling, identification and analysis of the TCA cycle (tricarboxylic acid cycle, citric acid cycle) and the ETC (electron transport chains, respiratory chains) of Escherichia coli. One of the SysMO-DB pals for SUMO. Now: Industrial affiliation
Projects: SysMO-LAB
Institutions: Heidelberg Institute for Theoretical Studies
Projects: MRRDEP
Institutions: Technische Universität Berlin (TU Berlin)
 https://orcid.org/0000-0002-1844-1168
https://orcid.org/0000-0002-1844-1168
Projects: COVID-19 Disease Map
Institutions: Paris Brain Institute / Inria
Projects: MESI-STRAT
Institutions: University of Innsbruck
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Tübingen
Trained as a Computer Scientist at the University of Jena, Germany, my interests drifted more and more towards "computing life" during my PhD at Simon Fraser University and the University of British Columbia, Canada. On one hand, that term captures that I got fascinated by the idea to understand life as a form of computation and to describe, reprogram, and reassemble parts of cells as described in many brilliant experiments from a spectrum of disciplines ranging from DNA computing to Synthetic ...
Projects: SysMO-LAB
Institutions: University of Rostock
Projects: Test Project
Institutions: Howest University of Applied Sciences
 https://orcid.org/0000-0002-9310-1876
https://orcid.org/0000-0002-9310-1876
Projects: COVID-19 Disease Map, ModeleXchange initiative
Institutions: The European Bioinformatics Institute < EMBL-EBI
Projects: TRALAMINOL
Institutions: CSIC Barcelona - Consejo Superior de Investigaciones Científicas
Projects: SysMetEx
Institutions: Systems Biomedicine Center, University of Luxembourg
Projects: Escherichia coli chemical product tolerance
Institutions: Abengoa Research SL
 https://orcid.org/0000-0003-2377-9929
https://orcid.org/0000-0003-2377-9929
Projects: DigiSal-BT8121
Institutions: Norwegian University of Science and Technology
Projects: TRANSLUCENT
Institutions: University of Leiden
Projects: MetApp, C1Pro, FUNCEMM
Institutions: INSA Toulouse, INRAE
 https://orcid.org/0000-0003-1312-3002
https://orcid.org/0000-0003-1312-3002
Dr. Stephanie Heux is leader of the MetaSys Team at LISB of Toulouse. She has 14 years of experience with microorganism as a model, metabolic engineering and NMR & MS-based metabolomics and fluxomics. After a 2-year Marie Curie fellowship at ETH Zurich where she developed advanced metabolomics methods and data mining approaches, she joined the LISBP as INRA researcher to undertake systems biotechnology research. She is interested in the microbial metabolism and, in particular, its organization, ...
Projects: CC-TOP
Institutions: Autonomous University of Madrid
 https://orcid.org/0000-0001-5740-5584
https://orcid.org/0000-0001-5740-5584
Projects: iRhythmics
Institutions: University of Rostock
Projects: BaCell-SysMO
Institutions: University of Erlangen-Nuernberg
CURRICULUM VITAE
Wolfgang Hillen Full Professor and Chairman of Microbiology Lehrstuhl für Mikrobiologie Institut für Biologie Friedrich-Alexander Universität Erlangen-Nürnberg Staudtstr. 5 91058 Erlangen, Fed. Rep. of Germany
Date of Birth: April 24, 1948 Place of Birth: Osnabrück, FRG Nationality: German Status: Married Children: Hauke Sven Hillen - May 17, 1987
RESEARCH AND PROFESSIONAL EXPERIENCE
Present Full Professor and Chairman of Microbiology at the Institute of Biology, Friedrich-Alexander ...
Projects: SYSTERACT
Institutions: University of Tübingen
Projects: AquaHealth (ERA-BlueBio)
Institutions: Sea & Sun Technology GmbH
Projects: Rhodolive
Institutions: National Institute of Chemistry (Slovenia)
Projects: STREAM
Institutions: University of Warwick
Projects: ILS Ceramide Ring Trial
Institutions: Forschungszentrum Jülich
 https://orcid.org/0000-0002-6540-6875
https://orcid.org/0000-0002-6540-6875
Projects: BaCell-SysMO
Institutions: University of Marburg
Projects: iRhythmics
Institutions: Leibniz Institute for Farm Animal Biology (FBN)
 https://orcid.org/0000-0003-2018-2836
https://orcid.org/0000-0003-2018-2836
Projects: AquaHealth (ERA-BlueBio)
Institutions: Hamburg University of Technology
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Gothenburg
Projects: SUMO
Institutions: University of Sheffield
Professor of Computer Science, University of Sheffield. FBCS, FIMA, CEng, C.Math, CITP. I have been involved in the use of computational techniques for modelling biological systems since 1980. More recently I have developed a technique of agent-based modelling based on the framework FLAME which is the only such system that can be run on supercomputers. We have made significant new biological discoveries using this approach: The approach models the location and activity of millions of individual ...
Projects: NTNU Health Druglogics
Institutions: Norwegian University of Science and Technology
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Projects: COSMIC
Institutions: Technical University of Munich
Projects: COVID-19 Disease Map
Institutions: Morpho-Biomimicry research and innovation centre
Currently I focuse on the integration of data into multi-scale models with statistical methods and uncertainty tracking in the research unit QuaLiPerF.
Projects: Not specified
Institutions: Not specified
STEMart is an industry-leading eCommerce platform incorporated with an extensive global footprint and a broad portfolio of more than 10,000 products. It aims to provide better lab materials, medical instruments and consumables, excellent technologies, and high-quality services to global customers in the fields of science, technology, and engineering, from the discovery stage upward to the manufacturing process. STEMart is dedicated to enhancing research and biotech ...
Projects: FAIRDOM user meeting
Institutions: Caltech
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: TRANSLUCENT
Institutions: University of Vienna
Projects: NL-Bioimaging FAIR Metadata Templates
Institutions: University of Leiden
 https://orcid.org/0000-0002-3704-3675
https://orcid.org/0000-0002-3704-3675
Projects: NAMPT affinity, NAD phylogeny
Institutions: University of Bergen
Projects: COVID-19 Disease Map
Institutions: Università Cattolica del Sacro Cuore
Projects: NAD COMPARTMENTATION
Institutions: University of Bergen
Projects: COVID-19 Disease Map
Institutions: University Maastricht
 https://orcid.org/0000-0002-8416-4533
https://orcid.org/0000-0002-8416-4533
Projects: FAIRDOM, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, LiSyM Core Infrastructure and Management (LiSyM-PD), ModeleXchange initiative, SDBV/HITS, EDITH (Ecosystem Digital Twins in Health) test project
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-8318-3222
https://orcid.org/0000-0002-8318-3222
Projects: Thesis
Institutions: CSIC Madrid
Projects: SysMO-LAB, Sustainable co-production
Institutions: University of Amsterdam, Wageningen University & Research
Projects: COVID-19 Disease Map
Institutions: Auckland Bioengineering Institute
Projects: Light control of leaf development
Institutions: University of Edinburgh
Projects: DigiSal, GenoSysFat
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6097-2539
https://orcid.org/0000-0001-6097-2539
Projects: Systems toxicology of Atlantic cod
Institutions: University of Oslo
 https://orcid.org/0000-0001-5453-2932
https://orcid.org/0000-0001-5453-2932
Projects: COVID-19 Disease Map
Institutions: University of Rome Tor Vergata
Projects: Cell4Chem, ODDITY, WATCHER, VIALACTEA
Institutions: Universidade de Santiago de Compostela
 https://orcid.org/0000-0003-3948-8077
https://orcid.org/0000-0003-3948-8077
Projects: KOSMOBAC
Institutions: Max-Planck-Institute of Biochemistry Martinsried
The focus of our research is the protein biogenesis and how stress-related factors modulate it. Protein biogenesis in general comprises various processes, i.e., translation, protein folding, each of which responds differently to external stress stimuli. Using systems biology approaches we seek to understand the interplay between these processes in fine-tuning the protein pattern and proteins’ abundance under osmotic stress conditions.
Projects: SysMilk
Institutions: Chalmers University of Technology
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
I am a PostDoc in prof. Brautaset's lab at NTNU in Trondheim, Norway. During my Ph studies D, Irla was involved in two ERA projects; SynMet and MetAPP (PAL) and currently in I am active in C1Pro project (Asset housekeeper). I have been working with methylotrophic Bacillus methanolicus since 2012, in that time I have been involved in engineering of that bacterium for production of different value-added products (amino acids and their derivatives, vitamins, and others). Furthermore, I have improved ...
Projects: COVID-19 Disease Map
Institutions: Zanjan University of Medical Sciences
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Okayama University
A (Cand) Doctor from Environmental Science at Padjadjaran University. My research is usually about ecosystem services in freshwater especially in the fisheries aspect of provisioning services.
Projects: Group Data Science, Methylation differences during respiratory infection, Implementation of Nanopore Sequencing for Detection of Treatment Induced Transcriptomic and Epitranscriptomic Changes in Leukaemic Tumour Models, OMICS of vascular smooth muscle cells under oxidative stress, Systems genetics of lung health and asthma (GWAS, PGS, QTL), iMED-RISK: Improvement of risk stratification in infant medulloblastoma using integrated models of clinical, histopathological and molecular data
Institutions: Forschungszentrum Borstel
Projects: COVID-19 Disease Map
Institutions: University of Heidelberg
Projects: COSMIC
Institutions: University of Nottingham
I am a Birmingham and MRC Fellow in mathematical biology. Specialising in the modelling of gene regulation networks using both numerical and analytical approaches, my work spans a range of biological applications, from drug development to bioenergy to understanding bacterial behaviour. My MRC fellowship gave me the opportunity to gain experimental training in order to generate the complementary data required to adopt a truly interdisciplinary approach to mathematical modelling in biology.
I worked ...
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur
Projects: PSYSMO
Institutions: University of Braunschweig
Projects: PSYSMO
Institutions: University of Duesseldorf
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University Medical Centre Groningen
Projects: PSYSMO
Institutions: University of Braunschweig
Projects: EmPowerPutida
Institutions: BASF
Projects: SulfoSys - Biotec
Institutions: University Bielefeld
Projects: FAIRDOM user meeting
Institutions: KAIST
Projects: STREAM
Institutions: University of Groningen
Projects: COSMIC
Institutions: University of Rostock
PhD University of Rostock, Germany Institute of Biological Sciences Division of Microbiology Albert-Einstein-Str. 3 18051 Rostock
Projects: C19DM-Neo4j
Institutions: Luxembourg Centre for Systems Biomedicine (LCSB)
 https://orcid.org/0000-0003-2047-0897
https://orcid.org/0000-0003-2047-0897
Projects: COVID-19 Disease Map
Institutions: American University of Sharjah
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
Projects: BESTER
Institutions: Green Biologics Ltd
Projects: SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, VHP project
Institutions: University Maastricht
Projects: Testing FairdomHub functionality
Institutions: University of Connecticut
Projects: SysMilk
Institutions: Chalmers University of Technology
Projects: MIX-UP
Institutions: RWTH Aachen University
Projects: SUMO
Institutions: University of Sheffield
Projects: Not specified
Institutions: Not specified
My Ph. D. project is connected to the project “Mørke flekker I laksefilet: Årsak til dannelse og titak som hemmer utvikling (EX-SPOT)”. The project includes small and commercial-scale experiments with high industry involvement to evaluate the influence of the environment and diet in developing discolored spots. Additionally, targeted studies are conducted to improve the knowledge of fundamental mechanisms through transcriptomic and sequencing.
Projects: GenoSysFat, DigiSal, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-5597-8397
https://orcid.org/0000-0001-5597-8397
Projects: iRhythmics
Institutions: University Medicine of Greifswald
Projects: Not specified
Institutions: Not specified
Hey! I am Leila Johnson and I will be your helper in academic writing. My passion to writing has revolved into the everyday work at EssaysWorld.net https://essaysworld.net/personal-statement-writing-service and now I am able not only to write just simply for myself, but to help those who are in need of any academic writing assistance. Stay with me and will teach you essential writing tips.
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Åbo Akademi University
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0003-4110-0748
https://orcid.org/0000-0003-4110-0748
Projects: Stress granules
Institutions: University of Bergen
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
 https://orcid.org/0000-0001-9318-4264
https://orcid.org/0000-0001-9318-4264
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Projects: FAIRDOM user meeting
Institutions: KAIST
Projects: TRANSLUCENT
Institutions: University of Cordoba
Projects: Hi-IMPAcTB, MIT SRP, MetNet, Cancer Systems Biology Consortium (CSBC), Endometriosis
Institutions: Massachusetts Institute of Technology
Projects: AquaHealth (ERA-BlueBio)
Institutions: Department of Planning, Aalborg University
Projects: CoolWine
Institutions: VTT Technical Research Centre of Finland
Projects: BlueTools Test
Institutions: FZJ
Projects: KOSMOBAC
Institutions: Ludwig-Maximilians University of Munich
I have the chair for Microbiology at the Ludwig-Maximilians-Universität München since 2004. My lab has a long-standing interest in stress response and transmembrane signal transduction in E. coli and V. harveyi. Our work is focused on the molecular mechanisms of membrane-integrated receptors, and the systems biological analysis of regulatory circuits.
Projects: SysVirDrug
Institutions: Technische Universität Dresden
Projects: Data Repository
Institutions: University of Ulm
Projects: TRANSLUCENT
Institutions: University of Applied Sciences Koblenz, Rhein Ahr Campus
Grammar school until 1998 1998-99 Alternative civilian service 1999-2002 Professional education as male nurse 2002-2007 Study of Biomathematics 2007-... PhD student in TRANSLUCENT project
Projects: HUMET Startup
Institutions: University of Amsterdam
POSITION Professor of Experimental Neuroendocrinology at the Academic Medical Center (AMC) and the Hypothalamic Integration Mechanisms group at the Netherlands Institute for Neuroscience (NIN), Amsterdam, The Netherlands RESEARCH My major projects focus on understanding how the hypothalamic biological clock controls hormonal rhythms and energy metabolism. For this we mainly use the rat as an animal model. Important techniques include blood sampling from and targeted brain infusions in awake and ...
MSD Animal Health: Boxmeer, Netherlands (Bioprocess Technology & Support)
Projects: Auromega
Institutions: Norwegian University of Science and Technology
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: SysMO-LAB, de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases
Institutions: Heidelberg Institute for Theoretical Studies
I'm a biologist working in the field of scientific datases as a biocurator.
Projects: Rhodolive, COVID-19 Disease Map
Institutions: Düzen Biological Sciences R&D and Production Company
Projects: Rhodolive
Institutions: Düzen Biological Sciences R&D and Production Company
Projects: BioZEment 2.0
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-6626-8697
https://orcid.org/0000-0002-6626-8697
Projects: SAFE-Aqua
Institutions: National Center for Genetic Engineering and Biotechnology
Projects: Simulation Foundries
Institutions: University of Stuttgart
Alexander Kel received his Ph.D. in Bioinformatics, Molecular Biology and Genetics in 1990. He studied biology and mathematics at Novosibirsk State University and obtained his M.S. in biology with special focus on mathematical biology in 1985. He worked for 15 years at the Institute of Cytology and Genetics, Russia (ICG) holding positions as a programmer, scientist, senior scientist and Vice-Head of the Laboratory of Theoretical Molecular Genetics. In 1995, he won the Academician Belaev Award. ...
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Institute of Pathology
I am a postdoc at the Technical University in Munich. When I studied nutritional science I got more and more interested in molecular cancer research. My PhD thesis was in the field of molecular cancer research. In our lab we are interested in molecular basic research on cell culture level, especially regarding gastric cancer.
Projects: MESI-STRAT
Institutions: Neuroimmun GmbH
Projects: COSMIC
Institutions: Wageningen University & Research
I am assistant professor at the Laboratory of Microbiology and my interest is in the area of molecular microbiology. Research focuses on the analysis of the metabolism of anaerobic fermentative bacteria and archaea, especially with respect to biofuel production (hydrogen, butanol). Within SysMo our tasks concern the effect of butanol stress, using metabolomics and transcriptomics.
Projects: SAFE-Aqua
Institutions: Institut Pasteur
Projects: Pro-Inflammatory Response BLE by Mmm
Institutions: BNVL, National Agricultural Research and Development Institute
Projects: SilicoTryp, SYSTERACT, SynBio4Flav
Institutions: University of Glasgow, Chalmers University of Technology
 https://orcid.org/0000-0002-3593-5792
https://orcid.org/0000-0002-3593-5792
Projects: HUMET Startup
Institutions: Wageningen University & Research
 https://orcid.org/0000-0003-4488-7734
https://orcid.org/0000-0003-4488-7734
Dr. Sander Kersten received his PhD in Nutritional Biochemistry from Cornell University in 1997. After a postdoctoral stay in the laboratory of Dr. Walter Wahli at the University of Lausanne, Switzerland, he moved to Wageningen in 2000, initially as a fellow of the Royal Netherlands Academy of Arts and Sciences and later as Associate Professor. Since 2011 he is Full Professor in Molecular Nutrition and since 2014 chair of the Nutrition, Metabolism and Genomics group. His current research interests ...
Project manager of STRENDA and MIRAGE @Beilstein-Institut
Projects: WG1 - Compound libraries coordination and integration of compound design
Institutions: Mehmet Akif Ersoy University
Projects: Systems toxicology of Atlantic cod
Institutions: Norwegian University of Science and Technology
Projects: STREAM
Institutions: University of Warwick
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: SilicoTryp
Institutions: University of Glasgow
Projects: SysMilk
Institutions: European Molecular Biology Laboratory
Projects: COSMIC
Institutions: University of Nottingham
I am a mathematical modeller concerned primarily with applications in biology,
Projects: PSYSMO
Institutions: Imperial College London
Projects: CoolWine
Institutions: Norwegian University of Science and Technology
Projects: EbN1 Systems Biology
Institutions: Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures
Projects: DigiSal, GenoSysFat
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-7828-2309
https://orcid.org/0000-0001-7828-2309
Projects: COVID-19 Disease Map
Institutions: Systems Biology Institute (SBI)
Projects: MOSES
Institutions: University of Heidelberg
Projects: SynBio4Flav
Institutions: Chalmers University of Technology
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
Projects: MOSES
Institutions: Medical University Vienna
Projects: COVID-19 Disease Map
Institutions: Universität Konstanz
Projects: Cell4Chem
Institutions: Helmholtz Zentrum für Umweltforschung (UFZ)
 https://orcid.org/0000-0002-8643-340X
https://orcid.org/0000-0002-8643-340X
Projects: SulfoSys
Institutions: eGene Biotechnologie GmbH
Projects: FAIRDOM user meeting
Institutions: Centre for Digital Life Norway
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Institute of Cytology and Genetics
I am a biomodeler, PhD student. Actually, I've graduated from Novosibirsk State University on two specialities: my bachelor diploma is done in computer science and the master thesis is defended in information biology. So, I'm kind of drifting towards biology. I am a part of the Haploid Evolutionary Constructor project. Our research group studies are dedicated to the simulation of prokaryotic communities. Personally, I am involved into the simulation of spatially distributed bacterial communities ...
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: OXYMOD, INBioPharm
Institutions: SINTEF
Projects: TRANSLUCENT, HUMET Startup
Institutions: Humboldt-Universität zu Berlin, Humboldt University Berlin
Projects: PSYSMO
Institutions: University of Hannover
Projects: WP 2: Antimalarial drug encapsulation, Project Coordination, WP 5: Intradermal 3D drug device administration, WP 4: Evaluation of drug activity against multiple life cycle stages of malaria parasites, Unshackling Membrane Protein Research: New Amphiphilic Copolymers for Extraction of Stable, Active Membrane Proteins, WP1 - Structure-activity relationships of amphiphilic copolymers, WP2.1 - Mono-chain-end functional amphiphilic copolymers, WP2.2 - Hetero di-chain-end functional amphiphilic copolymers, WP2.4 - Block copolymers for tethering to surfaces, WP2.3 - Meta-stable amphiphilic copolymers, WP2.5 - Fusion of SMALPs, WP2.6 - SMALPs for photosynthetic research, WP3 - Scalability of polymer production process, WP4 - Community engagement and technical advice
Institutions: University of Stellenbosch, Stellenbosch University
 https://orcid.org/0000-0003-1561-274X
https://orcid.org/0000-0003-1561-274X
Projects: SUMO
Institutions: University of Stuttgart
I recently joined the Institute for System Dynamics as a PhD student. I'm currently working on a dynamic model of nitrate respiration in E. coli.
Projects: BaCell-SysMO
Institutions: University of Stuttgart
I am a biologist in the lab of Prof. Reuss at the University of Stuttgart and I am working in the field of biotechnology and mathematical modelling.
Projects: COVID-19 Disease Map
Institutions: Ankara University
 https://orcid.org/0000-0002-8347-3393
https://orcid.org/0000-0002-8347-3393
Projects: MESI-STRAT
Institutions: Medical University Innsbruck
Projects: DigiSal, GenoSysFat, SAFE-Aqua, Unlock
Institutions: Wageningen University & Research
 https://orcid.org/0000-0001-8172-8981
https://orcid.org/0000-0001-8172-8981
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: Lipofungi, DeepHyperSpec, LIGNOLIPP
Institutions: Norwegian University of Life Sciences
Projects: BaCell-SysMO
Institutions: University of Braunschweig
Projects: SulfoSys, FAIRDOM user meeting, Service to Milano-Bicocca with respect to their ATP-ROS model (Active NOW), Make Me My Model, Service to University of Lisbon (Portugal) with respect to their CFTR maturation model (Active NOW), Service to LCSB (Luxembourg) with respect to ROS management in Parkinson’s disease and cancer model (Active NOW), Service to URV Tarragona, Spain with respect to their Safety Assessment of Endocrine Disrupting Chemicals model (Active NOW), Service to Universidade Católica Portugues with respect to their Molecular Insight into Autism Spectrum Disorder (ASD) model (Active NOW), Service to Slovenia with respect to their Protease signaling network in neurodegeneration model (Active NOW), Service to University of Duisburg- Essen (Germany): with respect to their The Yin-Yang of Metabolism; Endometatoxicity (YYME) model (Active NOW), Service to Sheffield University (UK): with respect to Mitochondrial perfect adaptation model (Active NOW), Service to Sanquin (Amsterdam): with respect to Modelling of acute and chronic inflammation (Prospective), Service to Munich (Germany): with respect toCharged peptide to charged membrane binding model (Prospective), Training Hunfeld, EraCoBiotech 2 nd call proposal preparation, ROS detailed model for MSB manucript, Mechanism based modeling viral disease ( COVID-19 ) dynamics in human population, COVID-19 Disease Map, Modelling COVID-19 epidemics, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: University of Amsterdam, VU University Amsterdam, Infrastructure Systems Biology Europe, Systems Biomedicine Center, University of Luxembourg, Luxembourg Centre for Systems Biomedicine (LCSB)
I have modelling expertise in precise kinetic models of metabolism and signal transduction; metabolic control analysis, hierarchical regulation analysis, non-equilibrium dynamics, statistical mechanics, enzyme kinetics, flux balance analysis. Energy and carbohydrate metabolism in Archaea, Bacteria and human; ammonium assimilation in Bacteria; differential network-based drug design; cancer metabolic rewiring; cell cycle; genome wide metabolic map and inborn errors of metabolism; epigenetics.
Projects: EraCoBiotech 2 nd call proposal preparation
Institutions: geneXplain GmbH
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
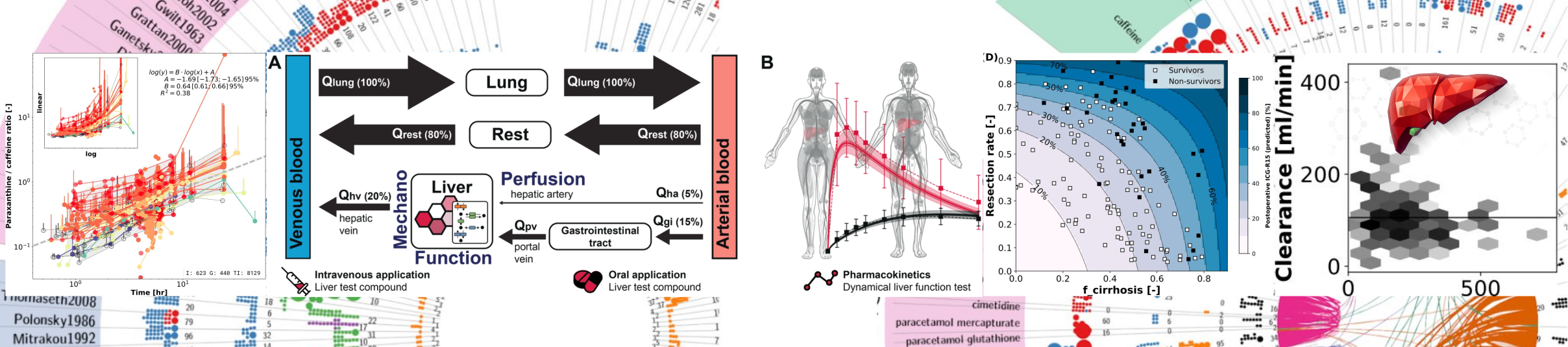
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: COVID-19 Disease Map
Institutions: Earlham Institute
Projects: FAIRDOM user meeting
Institutions: IDIBAPS
Projects: SulfoSys, SulfoSys - Biotec, Glucose metabolism in cancer cell lines
Institutions: University Duisburg-Essen, Stellenbosch University
Projects: PSYSMO
Institutions: Imperial College London
Projects: SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke
Institutions: University of Rostock
Projects: STREAM
Institutions: University of London UCL
Projects: HETCROP
Institutions: Institute of Plant Genetics, Polish Academy of Sciences
 https://orcid.org/0000-0001-5318-9896
https://orcid.org/0000-0001-5318-9896
Projects: icpm-kth
Institutions: KTH Royal Institute of Technology
 https://orcid.org/0000-0002-3828-6978
https://orcid.org/0000-0002-3828-6978
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Humboldt-Universität zu Berlin
Projects: ERNEST Mapping Group Pilot Study, COVID-19 Disease Map
Institutions: Humboldt-Universität zu Berlin
Projects: TRANSLUCENT
Institutions: Humboldt-Universität zu Berlin
I created this for all SysMo Modellers
http://www.semanticsbml.org/aym Annotate Your Model
There you can annotate your non SBML models with biological terms (MIRIAM annotations). As a cool extra you can view you model source code with inserted biological infomation.
Together with this http://www.semanticsbml.org/semanticSBML you can serach for similar BioModels. The similarity search is based on MIRIAM annotations that are attached to you model. AYM also allows you to create annotations without ...
Projects: COVID-19 Disease Map
Institutions: University College London (UCL)
 https://orcid.org/0000-0003-3171-6098
https://orcid.org/0000-0003-3171-6098
Projects: COSMIC
Institutions: University of Goettingen
I`m interested to investigate the Influence of the accumulation of reduction equivalents on solvent production
Projects: SilicoTryp
Institutions: University of Heidelberg
Professor of Biochemistry at the Centre of Biochemistry of Heidelberg University, teaching biochemistry for medical and biology students Research focus is the trypanothione redox metabolism of African trypanosomes (Trypanosoma brucei). The work is funded within the Collaborative Research Centre 544 on "Control of Tropical Infectious Diseases" of the German Research Foundation
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, ZucAt, SysMO-LAB, Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, ErasysApp Funders, EraCoBiotech 2 nd call proposal preparation, Service to URV Tarragona, Spain with respect to their Safety Assessment of Endocrine Disrupting Chemicals model (Active NOW), FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, Multiscale modelling of state transitions in the host-microbiome-brain network, BESTER, TRALAMINOL, Sustainable co-production, INDIE - Biotechnological production of sustainable indole, Extremophiles metabolsim, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, OLCIR: Optimization of Lung Cancer Therapy with Ionizing Radiation, NAD COMPARTMENTATION, HOTSOLUTE, Stress granules, FAIRDOM Community Workers, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", Mechanism based modeling viral disease ( COVID-19 ) dynamics in human population, COVID-19 Disease Map, AquaHealth (ERA-BlueBio), LiSyM Core Infrastructure and Management (LiSyM-PD), Early Metabolic Injury (LiSyM-EMI - Pillar I), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Modelling COVID-19 epidemics, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, SCyCode The Autotrophy-Heterotrophy Switch in Cyanobacteria: Coherent Decision-Making at Multiple Regulatory Layers, SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke, CC-TOP, BioCreative VII, MESI-STRAT Review, SDBV/HITS, MESI-Review 2024
Institutions: Heidelberg Institute for Theoretical Studies, FAIRDOM User meeting, Norwegian University of Science and Technology, University of Rostock, University of Innsbruck
 https://orcid.org/0000-0003-3540-0402
https://orcid.org/0000-0003-3540-0402
I am a researcher at the Scientific Databases and Visualization Group at Heidelberg Institute for Theoretical Studies (HITS) , one of the developers of SabioRK - System for the Analysis of Biochemical Pathways - Reaction Kinetics (http://sabiork.h-its.org/) . I am working on design and maintenance of the information systems to store, query and analyse systems biology data; definition and implementation of methods for the integration of data from multiple sources. In **[SySMO-DB ...
Projects: SysMO-LAB
Institutions: University of Rostock
PD Dr. rer. nat. habil. Bernd Kreikemeyer is the head of a research group at the Inst. of Medical Microbiology and Hospital Hygiene focusing on the pathogenesis of the human LAB Streptococcus pyogenes, microbial biofilm biology, and oral microbiology. Streptococcus pyogenes research, which is the relevant part for this BMBF proposal, is focused on 1. Identification of GAS virulence factors, 2. Mechanisms of GAS host cell adherence, host cell internalization, cytotoxicity of GAS towards host cells ...
I'm an engineer at the MPI Magdeburg and I'm working in the field of mathematical modeling, model verification, parameter identification, model analysis and experimental design. I'm involved in two projects, KOsmoBac and PSYSMO.
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur
Projects: COVID-19 Disease Map
Institutions: Centre national de la Recherche Scientifique (CNRS)
Projects: AquaHealth (ERA-BlueBio)
Institutions: University of Hamburg, Microbiology and Biotechnology
Projects: Virtual Human Platform for Safety Assessment
Institutions: Utrecht University
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: SulfoSys - Biotec
Institutions: University Duisburg-Essen
Projects: TRANSLUCENT
Institutions: University of Applied Sciences Koblenz, Rhein Ahr Campus
I am working in the mathematical modeling of potassium homeostasis. In addition, we are developing tools and methods for the statistical analysis of biological data.
Projects: SysMetEx
Institutions: TATAA BioCenter
Projects: MOSES
Institutions: Medical University Vienna
Projects: SYLOBIO
Institutions: Institut für Medizinische Mikrobiologie, Virologie und Hygiene
Projects: BaCell-SysMO
Institutions: University of Groningen
Group leader Molecular Genetics
Projects: COSMIC
Institutions: Wageningen University & Research
I'm an experimentalist 'Pre-doc' (I still have to finish my PhD thesis) and my work on the COSMIC project will focus on setting up a metabolomic analysis method for Clostridium acetobutylicum. In the past I have worked on metabolic engineering of the same organism by disrupting genes to asses their impact on acid and solvent formation. I'm looking forward to joining the COSMIC web-community. It hopefully will all us to stay in touch and update each other on advances in the (computer)lab.
Projects: INBioPharm
Institutions: Norwegian University of Science and Technology
Projects: COVID-19 Disease Map
Institutions: National Institute of Immunology
I am research scholar in the Department of Chemical Engineering at the University of Rovira i Virgili, Spain, member of the ENVIRONMENTAL ANALYSIS AND MANAGEMENT (AGA) group and also part of TECNATOX a specialized research center in the area of Technology Transfer in Toxicology, Food and Environmental Health.
MY RESEARCH INTERESTS:
Knowledge Engineering and System modelling in environmental domain. Uncertainty Modelling (Stochastic, Fuzzy and hybrid system) Integrated Modelling and decision ...
Projects: INBioPharm
Institutions: Norwegian University of Science and Technology
Projects: SysMO-LAB, Kinetics on the move - Workshop 2016, de.NBI-SysBio
Institutions: University of Heidelberg
Ursula Kummer is heading the dept. "Modeling of Biological Processes" at the University of Heidelberg.
Projects: FAIRDOM
Institutions: University of Zürich
Network biology, knowledge formalisation, cancer, data analysis
Projects: SysVirDrug
Institutions: Utrecht University
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation, HubMOL
Institutions: Mimetas, Mimetas (Netherlands)
Projects: COMBINE Multicellular Modelling
Institutions: University of St Andrews
Projects: GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University Medical Center Göttingen
 https://orcid.org/0000-0002-9895-2469
https://orcid.org/0000-0002-9895-2469
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: COVID-19 Disease Map
Institutions: University Maastricht
 https://orcid.org/0000-0002-7699-8191
https://orcid.org/0000-0002-7699-8191
Projects: Test project for Sciender
Institutions: Ural State Medical University
I hold a Medical Doctor Diploma (Lviv, Ukraine) with the specialization in General Medicine. After the graduation from the Post Graduate Program in Bioinformatics at the Seneca College/York University (Toronto, Canada), I successfully participated in the number of scientific projects conducted at the University of Toronto (Canada) and the Toronto East General Hospital (Canada).
I obtained the PhD in Bioinformatics at the Swiss Institute of Bioinformatics (Geneva, Switzerland). As a PhD student, ...
Projects: DigiSal, GenoSysFat
Institutions: Cargill
Projects: PhD project_ValeriaLaGatta
Institutions: Life Science, University of Modena and Reggio Emilia
Projects: HubMOL
Institutions: Ludwig-Maximilians University of Munich
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Imperial College London
I am a PhD student in the Theoretical Systems Biology group, based at Imperial College London. The aim of my PhD is to understand how noise can be the driving force of decision-making processes (differentiation, self-renewal, apoptosis or tumorgenesis), and what are our chances to control them. So far I have been working on method development for stochastic models, a moment closure framework and a stochastic reachability method, to look into cell-to-cell-variability.
Projects: Rhodolive
Institutions: Düzen Biological Sciences R&D and Production Company
Projects: Rhodolive
Institutions: Düzen Biological Sciences R&D and Production Company
Projects: BaCell-SysMO
Institutions: University of Greifswald
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
I am an engineer with a PhD degree in Chemical Engineering and had been working on dynamic modeling of mammalian cell culture fermentation in London for three years before moving into simulation of microbial systems. In this PSYSMO project I am mainly involved in modeling of PHAs synthesis. I am also a PAL since May 2009 - 2011 to coordinate data management and general communication among all 17 partners.
Projects: CropClock
Institutions: Systems Biomedicine Center, University of Luxembourg
Projects: MESI-STRAT
Institutions: University Medical Centre Groningen
Projects: CC-TOP
Institutions: University of Freiburg
I am a PhD student in the field of Systems Biology. In my PhD project I apply mathematical modelling to understand the role of time delay in biological systems containing delayed negative feedbacks.
Projects: FAIRDOM user meeting
Institutions: Institute of Cytology and Genetics SB RAS
Projects: LiVE
Institutions: University Medical Center Göttingen
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Professor- University Clermont Auyvergne (UCA) - Institute of Chemistry of Clermont-Ferrand (ICCF)
Projects: APPN Test Project
Institutions: Adelaide University
Projects: BioZEment 2.0, BESTER, Auromega, C1Pro, AquaHealth (ERA-BlueBio)
Institutions: SINTEF
Projects: NL-Bioimaging FAIR Metadata Templates
Institutions: University of Leiden
 https://orcid.org/0000-0002-0615-9616
https://orcid.org/0000-0002-0615-9616
Projects: COVID-19 Disease Map
Institutions: Swiss Institute of Bioinformatics
Projects: WP 3: Drug release kinetics study, WP 4: Evaluation of drug activity against multiple life cycle stages of malaria parasites, WP 5: Intradermal 3D drug device administration, WP 2: Antimalarial drug encapsulation, WP 1: Biodegradable and responsive polymer matrix, Project Coordination
Institutions: Leibniz-Institut für Polymerforschung Dresden e. V.
 https://orcid.org/0000-0002-1760-6426
https://orcid.org/0000-0002-1760-6426
Projects: SAFE-Aqua
Institutions: National Center for Genetic Engineering and Biotechnology
Projects: COVID-19 Disease Map
Institutions: University of California, Los Angeles
Projects: STREAM
Institutions: University of Warwick
Research fellow in Bioinformatics at the Warwick Systems Biology Centre, University of Warwick.
Working on high throughput data analysis (microarray data, next generation sequencing) and data integration (database management, text-mining, gene annotation via public databases)...
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Secretary
Projects: Not specified
Institutions: Not specified
Projects: TRANSLUCENT
Institutions: University of Applied Sciences Koblenz, Rhein Ahr Campus
Projects: CC-TOP
Institutions: University of Goettingen
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0002-0248-6679
https://orcid.org/0000-0002-0248-6679
Projects: Kinetics on the move - Workshop 2016
Institutions: Heidelberg Institute for Theoretical Studies
Research associate at HITS, software developer
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Xenophiles Systems Biology
Institutions: University of Amsterdam
Projects: SilicoTryp
Institutions: University of Heidelberg
Projects: SysMilk
Institutions: Biobyte Solutions GmbH (BS)
Projects: SysMO-LAB
Institutions: University of Heidelberg
I am working on a kinetic model of the central metabolism as well as on a genome wide model of Streptococcus pyogenes.
Projects: COVID-19 Disease Map
Institutions: Cardiff University
Projects: MIT SRP, Hi-IMPAcTB, MetNet
Institutions: Massachusetts Institute of Technology
 https://orcid.org/0000-0001-7363-562X
https://orcid.org/0000-0001-7363-562X
Projects: SYSTERACT, INBioPharm, OXYMOD, AquaHealth (ERA-BlueBio)
Institutions: SINTEF
Projects: BaCell-SysMO
Institutions: University of Newcastle
The main component of research in my group is in the determination of macromolecular structures by X-ray crystallography. However, since not all proteins crystallize, every effort is made to complete our understanding of how proteins function by utilizing other methods, such as microbial genetics, monitoring protein:ligand interactions by biochemical and biophysical methods, electron microscopy and bioinformatics
Projects: MESI-STRAT
Institutions: University of Groningen
MESI-STRAT Project Manager
Projects: Virtual Human Platform for Safety Assessment
Institutions: University Medical Center Utrecht
Projects: An overview of systematic reviews of treatment of polycystic ovary syndrome with Chinese medicine, Scalp Acupuncture and Treadmill Training Inhibits Neuronal Apoptosis Through Activating cIAP1 in Cerebral Ischemia Rats, Electroacupuncture preconditioning reduces oxidative stress in the acute phase of cerebral ischemia-reperfusion in rats by regulating iron metabolism pathways, The Effect of Nape Acupuncture Combined with Rehabilitation Training on Dysphagia After Stroke: A Meta-analysis, A Meta-Analysis of Functional Recovery of Aphasia after Stroke by Acupuncture Combined with Language Rehabilitation Training
Institutions: HLJUCM, Heilongjiang University of Chinese Medicine
 https://orcid.org/0000-0002-9184-5097
https://orcid.org/0000-0002-9184-5097
Projects: COVID-19 Disease Map
Institutions: University of Rome Tor Vergata
Projects: Hi-IMPAcTB, MetNet, MIT SRP
Institutions: MIT, Massachusetts Institute of Technology
Projects: TRANSLUCENT
Institutions: University of Bonn
Projects: XyloCut
Institutions: Lund University
Projects: BaCell-SysMO
Institutions: University of Rostock
Modelling of the general stress response activation cascade of sigB in B. subtilis in response to starvation.
Projects: SEEK tutorial for DigiSal
Institutions: Centre national de la Recherche Scientifique (CNRS)
In charge of educating on/proselytising data management practices at IFB, Institut Français de Bioinformatique https://www.france-bioinformatique.fr/.
Projects: HubMOL
Institutions: UiT The Arctic University of Norway
 https://orcid.org/0009-0003-7379-1771
https://orcid.org/0009-0003-7379-1771
Projects: SCaRAB
Institutions: Norwegian University of Science and Technology
Projects: Rhodolive
Institutions: National Institute of Chemistry (Slovenia)
I am had of the Research Group PiDOMICS which aims at the identification of human biomarkers for fungal infection using omics-data. Moreover, I am PI Infrastructure project of the Collaborative Research Center / Transregio 124 Pathogenic fungi and their human host: Networks of Interaction - FungiNet. Thereby my expertise is the implementation and usage of pipeline for OMICS (genome, transcriptome, protoem) data analysis, as well as data-warehouses for visualizing these data.
Projects: EmPowerPutida
Institutions: LifeGlimmer GmbH
Projects: ROBUSTYEAST
Institutions: Otto-von-Guericke University Magdeburg
 https://orcid.org/0000-0001-9899-2463
https://orcid.org/0000-0001-9899-2463
Projects: Rhodolive
Institutions: Leuphana University of Lüneburg
Projects: SCaRAB
Institutions: University of Greifswald
Projects: PSYSMO
Institutions: Imperial College London
Projects: CropClock
Institutions: Linköping University
Universitat Autónoma de Barcelona : Barcelona, Spain Dra. biotechnolog
Projects: GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University of Leipzig - Institute for Medical Informatics, Statistics and Epidemiology (IMISE)
 https://orcid.org/0000-0002-2344-0426
https://orcid.org/0000-0002-2344-0426
Projects: PhotoBoost
Institutions: University of Lancaster
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: EnzymeML
Institutions: University of Stuttgart
Projects: AquaHealth (ERA-BlueBio)
Institutions: Hamburg University of Technology
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Imperial College London
I am a first year PhD student at Imperial College London. My undergraduate degree was in Chemistry with a focus in medicinal Chemistry. My PhD project focuses on using chemical proteomics to obtain time resolved data of hypothesized signalling networks in cancer. More specifically, I study the roles of the KLK Activome in prostate cancer progression. My future research goals include developing a deterministic kinetic model of the KLK Activome that will aid in the discovery of novel therapeutics ...
Projects: Environmentally-friendly bioadhesives from renewable resources (WooBAdh), Ultreia, MPW-Val, Enzymatic biorecycling of PET (PETzyme), Integrated BIOfactory for the multiproduct valorisation of PIG MAnure (BIOPIGMA)
Institutions: Universidade de Santiago de Compostela
 https://orcid.org/0000-0003-4394-7432
https://orcid.org/0000-0003-4394-7432
Thelmo A. Lu-Chau is chemical engineer by the Universidad Nacional de Trujillo (Perú) and PhD in Chemical Engineering by the Universidade de Santiago de Compostela (USC, Spain). He is currently working as Associate Professor at the Dept. Chemical Engineering at the Faculty of Sciences (Lugo) of the USC. His research areas include the production of biofuels (ethanol, butanol) and biopolymers (PHA) from lignocellulosic biomass and cereals, the production of enzymes (glucanases and ligninolytic ...
Projects: PhotoBoost
Institutions: KWS
Projects: TRANSLUCENT
Institutions: University of Bonn
Projects: BESTER
Institutions: PROCESSIUM
Projects: COVID-19 Disease Map, C19DM-Neo4j
Institutions: Harvard Medical School
 https://orcid.org/0000-0001-5709-371X
https://orcid.org/0000-0001-5709-371X
Projects: KOSMOBAC
Institutions: University of Aberdeen
I am a research technician at the Institute of Medical Science in Aberdeen, working for Prof. Ian Booth. The topic of our workpackage deals with K+ homostasis in Escherichia coli. I am working with the protein KefF, a regulatory subunit of the potassium channel KefC.
Projects: BaCell-SysMO
Institutions: University of Greifswald
I'm Post-Doc in the lab of Prof. Becher at the University of Greifswald. I'm working on the relative and absolute protein quantitation using gel-based and mass-spectrometric methods.
Associate Professor of Computational Biology at NTNU
Projects: BaCell-SysMO
Institutions: University of Greifswald
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: DeepHyperSpec
Institutions: Norwegian University of Life Sciences
Expert in the chemical sciences with specialisation in chemical synthesis and molecular logic-based computation.
Projects: COVID-19 Disease Map
Institutions: Biomax Informatics AG
Projects: SUMO
Institutions: University of Sheffield
I am a research associate in the department of computer science at the University of Sheffield since January 2008. My research is primarily involved with using agent-based modelling techniques and mathematical modelling techniques to model Escherichia coli K-12 Respiratory Adaptation. My research interests also include, development of workflows to analyze Microarray Data.
Projects: B.Ed Colleges in Bangalore
Institutions: B.Ed Colleges in Bangalore
Projects: Munich Cluster for Systems Neurology
Institutions: University of Munich LMU
 https://orcid.org/0000-0001-9212-2520
https://orcid.org/0000-0001-9212-2520
Projects: OXYMOD
Institutions: Vectron Biosolutions AS
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
Projects: SulfoSys
Institutions: University of Vienna
Projects: PSYSMO
Institutions: Imperial College London
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
 https://orcid.org/0000-0001-5440-5503
https://orcid.org/0000-0001-5440-5503
PhD student
Projects: COVID-19 Disease Map
Institutions: Uppsala University
Projects: BaCell-SysMO
Institutions: University of Groningen
PhD student. Analyzing CcpA affinity to cre boxes (catabolite responsive elements) and response of B. subtilis to membrane protein overproduction stress.
Projects: SysMilk
Institutions: Chalmers University of Technology
Projects: XyloCut
Institutions: Forschungszentrum Jülich
Projects: FAIRDOM user meeting
Institutions: IDAEA-CSIC
Projects: TRALAMINOL
Institutions: CSIC Barcelona - Consejo Superior de Investigaciones Científicas
Projects: COVID-19 Disease Map
Institutions: Centre de Recherche en Cancerologie de Toulouse
Projects: BESTER, BioZEment 2.0
Institutions: SINTEF
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0002-5914-2578
https://orcid.org/0000-0002-5914-2578
Projects: ComPASs - Common Pathways in Amyotrophic Lateral Sclerosis (ALS) and Spinal Muscular Atrophy (SMA), miRiAD - exploring the role of microRNAs in T cell function and anti-viral defence, LungCARD - Blood test for clinical therapy guidance of non-small cell lung cancer patients
Institutions: RNA Systems Biology Lab - BioISI/FCUL
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: BaCell-SysMO
Institutions: University of Groningen
Projects: ERNEST Mapping Group Pilot Study
Institutions: MRC Laboratory of Molecular Biology
 https://orcid.org/0000-0003-0373-8927
https://orcid.org/0000-0003-0373-8927
Projects: STREAM
Institutions: INBIOTEC Leon
Projects: EmPowerPutida
Institutions: Wageningen University & Research
This profile is created to handle PoLiMeR data only - mainly the materials from Anne-Claire M.F. Martines PhD thesis (temporary solution)
Projects: CC-TOP
Institutions: Autonomous University of Madrid
Projects: EmPowerPutida
Institutions: CSIC
Projects: Dynamic interplay between nuclei and mitochondria in ageing cells
Institutions: University of Newcastle
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
 https://orcid.org/0009-0004-0159-7539
https://orcid.org/0009-0004-0159-7539
Projects: PSYSMO, DigiSal, GenoSysFat, HUMET Startup, EmPowerPutida, MycoSynVac - Engineering Mycoplasma pneumoniae as a broad-spectrum animal vaccine, SAFE-Aqua, INDIE - Biotechnological production of sustainable indole
Institutions: Helmholtz Centre for Infection Research Braunschweig, Wageningen University & Research
My research activities has been to use mathematical models and Computational Biology to answer biological questions, intertwining in silico and experimental methods at all stages. I have a strong interest in exploring the interfaces between Fundamental Biology and bona fide Engineering, specifically in the realm of environmental and industrial problems. The research goals of my group are to contribute to the elucidation of mechanisms underlying basic cellular processes, evolution and ecological ...
Projects: WineSys
Institutions: Norwegian University of Science and Technology
Projects: COVID-19 Disease Map
Institutions: Accelera
Institutions: Universitat Rovira i Virgili
Projects: DigiSal-BT8121
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0001-5411-8969
https://orcid.org/0000-0001-5411-8969
I am part of 180 ̊ N Project: Clinical PET/MRI in collaboration with St Olav’s Hospital. In Work Package-5, we aim to develop and validate new machine learning methods that exploit the quantitative nature of the PET and MR images. We will develop a pipeline for pre-processing and quality assurance of the acquired images, as well as a toolbox with machine learning algorithms focusing on region classification and detection of change over time. These methods will be applied to answer open clinical ...
Projects: COVID-19 Disease Map
Institutions: Swiss Institute of Bioinformatics
Projects: SilicoTryp
Institutions: University of Edinburgh
Projects: SilicoTryp
Institutions: University of Edinburgh
Projects: COVID-19 Disease Map
Institutions: NYU Grossman School of Medicine
Projects: Cell4Chem, ODDITY, WATCHER, VIALACTEA, AQUACIRCLE, FATCRACK
Institutions: Universidade de Santiago de Compostela
 https://orcid.org/0000-0001-8766-8908
https://orcid.org/0000-0001-8766-8908
Projects: PSYSMO
Institutions: University of Essex
Wet Lab experimentalist. Solute stress in Pseudomonas putida, group 12 in the PSYSMO consortium under Terry McGenity at the University of Essex. A PSYSMO PAL.
Projects: PSYSMO
Institutions: University of Essex
Projects: COMBINE Multicellular Modelling
Institutions: University of Newcastle-upon-Tyne
 https://orcid.org/0000-0002-8361-2795
https://orcid.org/0000-0002-8361-2795
Interdisciplinary researcher; passionate about findable, accessible, interoperable, and reproducible (FAIR) scientific knowledge.
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Post doc. in the SysMO-LAB2 project from August 2010. I work at Nofima and the Norwegian University of Life Sciences (UMB) at Ås, Norway. My focus in SysMO-LAB2 will be on four Lactobacillus plantarum strains, diversity analysis, omics-technologies, genome scale modelling.
Background: Ph.D. in Molecular Microbiology June 2010, where I worked with Lactobacillus sakei, metabolism and diversity studies.
Projects: ZucAt
Institutions: Novosibirsk State University
I got my Master’s Degree in Biomathematics, Bioinformatics, and Computational Biology 2007 at Novosibirsk State University (NSU). I am working at Computer Proteomics Laboratory, Institute of Cytology and Genetics SB RAS. We develope computer system to analyze the coding features of functional sites by taking into account the exon structure of the gene, to detect the exons involved in shuffling in protein evolution, also to design protein-engineering experiments. Tools developed - SitEx http://www-bionet ...
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
 https://orcid.org/0000-0001-6142-5270
https://orcid.org/0000-0001-6142-5270
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Projects: RootBook
Institutions: University of Freiburg
Projects: COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University of Leipzig - Institute for Medical Informatics, Statistics and Epidemiology (IMISE)
 https://orcid.org/0000-0002-9256-7543
https://orcid.org/0000-0002-9256-7543
Projects: Group Data Science
Institutions: Forschungszentrum Borstel
Projects: COVID-19 Disease Map
Institutions: Monash University
Projects: MOSES
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: SEEK tutorial for DigiSal, DigiSal
Institutions: Norwegian University of Life Sciences
Projects: MOSES
Institutions: VU University Amsterdam
Projects: STREAM
Institutions: University of Groningen
Projects: MOSES
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: Unlock
Institutions: Wageningen University & Research
Projects: COVID-19 Disease Map
Institutions: Estonian Genome Centre, Institute of Genomics, University of Tartu
Projects: MetApp
Institutions: ETH Zurich
Projects: BaCell-SysMO
Institutions: University of Greifswald
Projects: BaCell-SysMO
Institutions: University of Goettingen
Projects: Not specified
Institutions: Not specified
BalaJi MicroTechnologies (BMT) is New Delhi, India based company, We are registered in India under Indian companies’ act 1955. We are Multi-Million USD turnover company & unit of "B.B. Group of Companies". The group is into business since 1949.
"BalaJi MicroTechnologies (BMT) is India's No.1 manufacturer of high performance Machine Vision & Line Scan Cameras & precision Machine Vision lenses under our brand name "BMT" which is also known as "BMT Machine Vision lens ". Our company core ...
Projects: COVID-19 Disease Map
Institutions: Policlinico Universitario Agostino Gemelli IRCCS
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Projects: KOSMOBAC
Institutions: University of Groningen
Kosmobac, WP3, looking at diffusion of macromolecules in vivo (in E.coli cells) and cell responses to osmotic shock using confocal (fluorescence) microscopy especially pulsed - FRAP
Projects: CC-TOP
Institutions: Faculty of Chemical Engineering and Technology
 https://orcid.org/0000-0003-0712-2417
https://orcid.org/0000-0003-0712-2417
Projects: BIDS
Institutions: Medizinische Hochschule Hannover
Projects: Millar group, TiMet, PHYTOCAL: Phytochrome Control of Resource Allocation and Growth in Arabidopsis and in Brassicaceae crops, POP - the Parameter Optimisation Problem, Regulation of flowering time in natural conditions, PlaSMo model repository
Institutions: University of Edinburgh
 https://orcid.org/0000-0003-1756-3654
https://orcid.org/0000-0003-1756-3654
Projects: COSMIC, BaCell-SysMO
Institutions: University of Rostock
Modelling of cellular signalling, Dynamic Motifs and Feedback, Quantitative Measures, Theoretical Aspects of Modelling Biological Systems
Projects: Not specified
Institutions: Not specified
Lifeasible provides a wide range of Yeast Competent Cells for research use.
Projects: KOSMOBAC
Institutions: University of Aberdeen
I am a biologist by training. My research career is focussed on the structure-function relationships of membrane transport systems in Escherichia coli. I am interested in understanding the mechanism of ion and solute transport across the membrane and how this influences bacterial cell survival. My work mainly focusses on the ligand-gated potassium efflux systems which are crucial for cell survival during electrophile exposure and the mechanosensitive channels involved in hypoosmotic stress ...
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
Projects: INDIE - Biotechnological production of sustainable indole
Institutions: Wageningen University & Research
Projects: COSMIC
Institutions: University of Nottingham
A molecular microbiologist with a passion for Clostridia! Interested in the development of more effective countermeasures (diagnosis, prevention & treatment) against pathogens, specifically Clostridium difficile and Clostridium botulinum as well as the exploitation of the medical and industrial properties of beneficial strains, specifically in cancer therapy and biofuel production
Projects: COVID-19 Disease Map
Institutions: University of Barcelona
Projects: BioRECIPE representation format
Institutions: University of Pittsburgh
 https://orcid.org/0000-0003-1110-3403
https://orcid.org/0000-0003-1110-3403
Projects: ROBUSTYEAST
Institutions: Ecole Polytechnique Fédérale de Lausanne (EPFL)
 https://orcid.org/0000-0001-7333-8211
https://orcid.org/0000-0001-7333-8211
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: COVID-19 Disease Map
Institutions: Earlham Institute
 https://orcid.org/0000-0001-9412-6867
https://orcid.org/0000-0001-9412-6867
Projects: GWI-HDMT stemcells
Institutions: SURF
Projects: COVID-19 Disease Map
Institutions: Golestan University of Medical Sciences
Projects: HETCROP
Institutions: Institute of Plant Genetics, Polish Academy of Sciences
 https://orcid.org/0000-0002-4512-845X
https://orcid.org/0000-0002-4512-845X
Projects: Not specified
Institutions: Not specified
I finished my biotechnology studies at Pablo de Olavide University (Seville) in 2010 and after that got a master degree in microbiology (Granada University). During my PhD (2010-2014) at Estación Experimental del Zaidín (Granada, Spain), I focused on the study of the biosynthesis of antibiotic compounds and the signaling mechanisms underlying antibiotic resistance in Pseudomonas putida from a transcriptomic and molecular point of view.
Projects: NIRBTEST
Institutions: Amsterdam UMC
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Uni Basel
Projects: COVID-19 Disease Map
Institutions: Institut Curie
Projects: MESI-STRAT
Institutions: Vall d´Hebron Institute of Oncology (VHIO)
Projects: COVID-19 Disease Map
Institutions: Barcelona Supercomputing Center
 https://orcid.org/0000-0002-7696-1241
https://orcid.org/0000-0002-7696-1241
I completed my BSc in Biology and MSc in Cell Biology by the University of Valencia. During my last undergrad year I participated in synthetic biology’s iGEM competition where I dove in the use of models in Biology, which pushed me to pursue a PhD in the Department of Applied Mathematics in the Technical University of Valencia.
My research on Metabolic Engineering of hydrogen in cyanobacteria led me to be visiting researcher at Uppsala University, Denmark Technical University and EMBL Heidelberg. ...
Projects: COVID-19 Disease Map
Institutions: Barcelona Supercomputing Center
Projects: HOTSOLUTE
Institutions: Consiglio Nazionale delle Ricerche
 https://orcid.org/0000-0002-3399-7973
https://orcid.org/0000-0002-3399-7973
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München / Institute of Experimental Genetics
Projects: STREAM
Institutions: University of Warwick
Systems Biologist specialising in data integration, high-throughput sequence analysis, and evolutionary and comparative analyses.
Projects: TRALAMINOL
Institutions: CSIC Barcelona - Consejo Superior de Investigaciones Científicas
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Stockholm University
Projects: EmPowerPutida
Institutions: LifeGlimmer GmbH
Computational Biologist
Projects: Fluid flow project
Institutions: Institute of Physiology Academy of Sciences
Projects: KOSMOBAC
Institutions: University of Aberdeen
Projects: FAIRDOM
Institutions: University of Manchester - Department of Computer Science
 https://orcid.org/0000-0003-1604-1512
https://orcid.org/0000-0003-1604-1512
FAIRDOM Project Wrangler
Associate Professor at the University of Santiago de Compostela. Expertise on biological urban and industrial wastewater treatment, nitrogen removal via conventional nitrification and denitrification processes and anammox, operation of aerobic granular sludge reactors and biopolymers production from industrial wastewater.
Projects: KOSMOBAC
Institutions: University of Aberdeen
Physicist, working on the modelling side.
Projects: TRANSLUCENT
Institutions: University of Bonn
Projects: HOTSOLUTE
Institutions: Evonik Industries AG
Projects: Systo models, PlaSMo model repository, Agro-ecological modelling
Institutions: University of Edinburgh
Projects: Hi-IMPAcTB
Institutions: Massachusetts Institute of Technology
 https://orcid.org/0000-0002-3455-2992
https://orcid.org/0000-0002-3455-2992
Projects: Xenophiles Systems Biology
Institutions: University of Amsterdam
Professor of microbial systems ecology at UvA Committee member PhD thesis Joost Aerts
Projects: WG1 - Compound libraries coordination and integration of compound design
Institutions: University of Münster
Projects: CC-TOP
Institutions: University of Freiburg
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, de.NBI-SysBio, Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, EnzymeML, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, MS_DILI, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COMBINE Multicellular Modelling, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, NMTrypI - New Medicines for Trypanosomatidic Infections, ModeleXchange initiative, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, BioCreative VII, SDBV ephemeral data exchanges, The BeeProject, SDBV/HITS, MESI-STRAT Review, MESI-Review 2024, DeepCurate, SABIO-VIS
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-4980-3512
https://orcid.org/0000-0002-4980-3512
I am group leader of the SDBV (Scientific Databases and Visualisation) group at the HITS gGmbH, the Heidelberg Institute for Theoretical Studies.
I am interested in finding data. Starting with my master's thesis I have always worked on how to store data in a way that you can find it, and how to make sense out of data that has been stored.
Within FAIRDOM I find interesting to help people to store their data in a way that they make sense even after years.
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur
Projects: DigiSal, GenoSysFat
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-0719-1308
https://orcid.org/0000-0003-0719-1308
Projects: iPlacenta- Placenta on a chip
Institutions: University of Dundee
 https://orcid.org/0000-0002-0274-819X
https://orcid.org/0000-0002-0274-819X
Projects: SysMO-LAB
Institutions: Wageningen University & Research
I'm a modeller, specialized in kinetic modeling of biochemical networks. My focus in the SysMO-LAB consortium is on creating models of Lactococcus lactis glycolysis and couple this to other related lactic acid bacteria like Streptococcus pyogenes and Enterococcus faecalis. Besides kinetic modeling, I'm also interested in combining various modeling techniques (genome-scale modeling, qualitative modeling).
Projects: BESTER
Institutions: Imperial College London
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
Projects: ErasysApp Funders
Institutions: Forschungszentrum Jülich
 https://orcid.org/0000-0002-9471-9328
https://orcid.org/0000-0002-9471-9328
Projects: COVID-19 Disease Map
Institutions: University of Bonn
Projects: BaCell-SysMO
Institutions: University of Greifswald
I am pursuing my PhD at Prof. Volker's Lab in the Department of Functional Genomics, EMA Universitat Greifswald, Germany. I am working on the general stress responses mediated by SigB and the prediction of SigB regulon members using the Random forest algorithm.
Projects: COVID-19 Disease Map
Institutions: University of Northampton
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Institut Curie
Projects: TRANSLUCENT
Institutions: University of Cordoba
PhD Student
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: COVID-19 Disease Map
Institutions: Riga Technical University
 https://orcid.org/0000-0003-2437-2055
https://orcid.org/0000-0003-2437-2055
Projects: CC-TOP
Institutions: CSIC Madrid
Project Manager at UAM
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Professor in biotechnology at the Dept. Chemistry, Biotechnology and Food Science. I am heading "Laboratory of microbial gene technology and food microbiology" that consists of approximately 20 members (staff members, technicians,and students). During the last 20 years my research has been focused on lactica acid bacteria with a focus on bacteriocins of lactic acid bacteria.These studies have included purification and chemical and genetic characterization of such peptides followed by biosynthesis ...
Projects: COVID-19 Disease Map
Institutions: Elsevier
Projects: LEANPROT
Institutions: Technische Universität Berlin (TU Berlin)
Projects: EbN1 Systems Biology, Adaptation of Salmonella enterica, Cross-Feeding B.theta, Pseudomonas aeruginosa and its vesicles, Phage infection in Nitrosomonas europaea
Institutions: Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures, Metabolomics and Services, Metabolomics and Services, Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures
Projects: SysMilk
Institutions: Chr-Hansen A/S (CH)
Projects: BaCell-SysMO
Institutions: University of Newcastle
I am a postdoc in the lab of Rick Lewis working on the biophysical characterization of protein-protein complexes involved in central carbon metabolism.
Software developer for FAIRDOM
I am an Associate Professor of Computational Systems Biology, at Univ Evry, University of Paris-Saclay. My research focuses on the application of computational systems biology approaches in human diseases, including the construction of disease maps, tool development for model inference, network integration and dynamical modelling.
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Milano-Bicocca
I am a PostDoc working on yeast metabolomics. During my PhD I studied the interplay between metabolism, cell cycle and signalling, mainly focusing on the Snf1/AMPK pathway. I am currently interested in studying metabolic rewiring caused by different nutrients, generating high-throughput data suitable for modelling.
Projects: Not specified
Institutions: Not specified
My name is Lola and I am a professional writer from papers-land where you can buy a powerpoint presentation, order professional essays, articles. Would be happy to help everyone in your beginnings. I really love my work and try to improve my writing skills each day. It is a dream work, so it brings me lots of positive emotions.
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: NTNU Health Druglogics, Colosys
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-6324-7524
https://orcid.org/0000-0002-6324-7524
Projects: SCaRAB, SysMilk, IMOMESIC
Institutions: Chalmers University of Technology
Projects: MESI-STRAT, NAD COMPARTMENTATION, Bergen(Ziegler lab) project AF-NADase
Institutions: University of Bergen
Institutions: University of Tuebingen, University of Tübingen
Projects: EmPowerPutida
Institutions: University of Stuttgart
Projects: SAFE-Aqua
Institutions: Wageningen University & Research
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: BaCell-SysMO
Institutions: University of Erlangen-Nuernberg
Projects: SCaRAB
Institutions: Norwegian University of Science and Technology
Projects: IMOMESIC
Institutions: Chalmers University of Technology
Projects: EmPowerPutida
Institutions: University of Stuttgart
PhD student, Institute of Biochemical Engineering at the University of Stuttgart
Projects: COSMIC
Institutions: University of Ulm
Postdoc University of Ulm/Dept. Microbiology and Biotechnology
Team leader "Quantitative Microbial Phenotyping" Institute of Bio- and Geosciences, IBG-1: Biotechnology Forschungszentrum Jülich GmbH 52425 Jülich, Germany
Projects: COVID-19 Disease Map
Institutions: Institut Curie
Projects: PhotoBoost
Institutions: Fraunhofer IME
Projects: PSYSMO, SynBio4Flav
Institutions: CSIC Madrid, CSIC
Projects: SUMO
Institutions: University of Stuttgart
Projects: DigiSal, GenoSysFat
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-1659-132X
https://orcid.org/0000-0003-1659-132X
Projects: SYSTERACT, INBioPharm
Institutions: SINTEF
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-5299-6450
https://orcid.org/0000-0001-5299-6450
Projects: MESI-STRAT
Institutions: PD-value B.V.
Projects: TRANSLUCENT
Institutions: University of Vienna
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Freiburg
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: CoolWine
Institutions: Norwegian University of Science and Technology
Projects: SUMO
Institutions: University of Sheffield
I am a first year PhD student, working with Professor Robert Poole (University of Sheffield), Professor Jeff Green (University of Sheffield) and Dr Jamie Wood (University of York) using a systems biology approach to study respiration in Escherichia coli.
Projects: SysMetEx
Institutions: TATAA BioCenter
Projects: RStudio-Shiny and Java_API connection testing, Fairdom Hub documents, Systems modelling age-related changes in the maintenance of dermal extra-cellular matrix: mechanisms and interventions, Outreach - Simulation of breast cancer development using an agent based modelling approach, Outreach - Breast Cancer RStudio-Shiny application, Outreach - Senescence RStudio-Shiny application, Outreach - Simulation of cellular senescence using an agent based modelling approach, Selective Destruction in Ageing
Institutions: University of Newcastle-upon-Tyne, University of Newcastle
 https://orcid.org/0000-0002-0107-711X
https://orcid.org/0000-0002-0107-711X
Projects: TRALAMINOL
Institutions: Université Clermont Auvergne (UCA)
I am a Researcher at the National Laboratory of Scientific Computing (LNCC). I am a bioinformatician and my research interests are in data integration and analyses. I use scientific workflows technologies to integrate and analyse data in systems biology, phylogenomics, and functional genomics.
Post Doc (2010 – 2015) of the Department of Computer Science at the COPPE Institute of the Federal University of Rio de Janeiro (Brazil) and was supported by a FAPERJ’s grant “Post-Doc Note 10” (2013 – ...
Projects: MetApp
Institutions: ETH Zurich
Projects: COVID-19 Disease Map
Institutions: National Institute of Informatics
 https://orcid.org/0000-0001-8725-3366
https://orcid.org/0000-0001-8725-3366
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Tokyo
I am a postdoc in The University of Tokyo. My research interest is to know structure of biological systems by functional relationships between genes working in response to endogenous and/or exogenous disruptions. I'm trying to find functional relationships between genes by similarity of phenotype defined with high-dimensional morphological features in yeast gene deletion collection. I'm happy if you will talk with me in any topics.
Projects: COVID-19 Disease Map
Institutions: Earlham Institute
 https://orcid.org/0000-0002-4903-6237
https://orcid.org/0000-0002-4903-6237
Biochemist currently keeping busy as: Research Data Manager (Vrije Universiteit Amsterdam), Software Engineer (Heidelberg University) and member of the SBML Development Team (Caltech).
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0002-5641-9420
https://orcid.org/0000-0002-5641-9420
Professor at Nord University, Bodø, Norway
Projects: STREAM
Institutions: University of Warwick
Postdoctoral Research fellow with experience in Genomics, transcriptomics, proteomics and metabolomics
Projects: MESI-STRAT
Institutions: German Cancer Research Center (DKFZ)
Clinical Trial Coordinator, Leader WP6 – Clinical Studies and Longitudal Analyses
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
Projects: ZucAt
Institutions: Institute of Cytology and Genetics, Novosibirsk State University
I am Senior Scientist at Institute of Cytology and Genetics SB RAS in Novosibirsk. My research focus is computational genomics: high-throughput sequencing, ChIP-seq, human genome research (HUGO member), high performance computing in bioinformatics
Projects: COVID-19 Disease Map
Institutions: UNIBI: Bielefeld University
 https://orcid.org/0000-0001-5977-798X
https://orcid.org/0000-0001-5977-798X
Projects: HUMET Startup
Institutions: AstraZeneca
Scientific Project Manager at Luxembourg Centre for Systems Biomedicine, University of Luxembourg http://lcsb.uni.lu
Projects: CropClock
Institutions: Systems Biomedicine Center, University of Luxembourg
Projects: COVID-19 Disease Map
Institutions: University of the Witwatersrand
Projects: COVID-19 Disease Map
Institutions: Centre for Research and Technology Hellas
Projects: COVID-19 Disease Map
Institutions: University of British Columbia
 https://orcid.org/0000-0001-5844-2731
https://orcid.org/0000-0001-5844-2731
Professor Christopher Overall was appointed a Tier 1 Canada Research Chair in Protease Proteomics and Systems Biology (2001) and an Honorary Professor Albert-Ludwigs Universität Freiburg, Germany (2014–) after being a Senior Fellow of the Freiburg Institute of Advanced Studies, Albert-Ludwigs Universität Freiburg, Germany (2010–2013). He was inducted as a fellow into the Royal Society of Canada, Academy of Science in 2018. He is best known for his development of proteomic methodology for the ...
Projects: COVID-19 Disease Map
Institutions: German Center for Neurodegenerative Diseases (DZNE)
Projects: COVID-19 Disease Map
Institutions: independent researcher
 https://orcid.org/0000-0002-3882-6073
https://orcid.org/0000-0002-3882-6073
Projects: WineSys
Institutions: Norwegian University of Science and Technology
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, GenoSysFat, DigiSal, FAIRDOM user meeting, FAIRDOM Community Workers
Institutions: University of Manchester - Department of Computer Science, Manchester Centre for Integrative Systems Biology, University of Manchester
 https://orcid.org/0000-0003-2130-0865
https://orcid.org/0000-0003-2130-0865
Senior Research Software Engineer and Architect working at the University of Manchester within the FAIRDOM team.
Leads the development of FAIRDOM SEEK and RightField.
Projects: COVID-19 Disease Map
Institutions: Pacific Northwest National Laboratory
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6990-0339
https://orcid.org/0000-0001-6990-0339
Projects: KOSMOBAC
Institutions: University of Aberdeen
I am final year PhD student in Prof Ian Booth's lab and a microbiologist by trade. I am interested in how enteric bacteria cope with stress and what systems they employ to increase their chances of survival, in particular upon methylglyoxal stress.
Projects: COMBINE Multicellular Modelling
Institutions: University of Heidelberg
 https://orcid.org/0000-0002-7637-7547
https://orcid.org/0000-0002-7637-7547
Projects: MESI-STRAT
Institutions: Vall d´Hebron Institute of Oncology (VHIO)
Projects: STREAM
Institutions: University of Groningen
Projects: FAIRDOM user meeting
Institutions: IDIBELL
Projects: WP 1: Biodegradable and responsive polymer matrix
Institutions: Leibniz-Institut für Polymerforschung Dresden e. V.
Projects: Project 3: Multipotent ABCB5(+) Stem Cells for Therapy of Age-related Skin Disorders, Project 1: Metabolic Reprogramming and Regeneration in the Aged Epidermis, Project 2: Epigenetic regulation of skin regeneration
Institutions: BWH, Harvard Chan School
 https://orcid.org/0000-0002-3859-3249
https://orcid.org/0000-0002-3859-3249
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: PhotoBoost
Institutions: University of Lancaster
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Gothenburg
Course participant
Projects: SysMO Funders
Institutions: BBSRC
Projects: Are microRNAs key mediators of cartilage destruction in osteoarthritis?
Institutions: University of Newcastle
Institutions: European Molecular Biology Laboratory
Projects: Differential abundance analysis of polyA segments in 3' sc/snRNAseq data
Institutions: University of Cologne
Assist. Prof. of Enzyme Technology Laboratory, Department of Chemistry, University of Crete, Greece
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: CC-TOP
Institutions: Hamburg University of Technology, Center for Free-Electron Laser Science (Uni Hamburg)
Projects: Systems modelling age-related changes in the maintenance of dermal extra-cellular matrix: mechanisms and interventions, Outreach - Senescence RStudio-Shiny application, Outreach - Simulation of cellular senescence using an agent based modelling approach
Institutions: University of Newcastle
 https://orcid.org/0000-0002-3792-6275
https://orcid.org/0000-0002-3792-6275
Projects: LEANPROT
Institutions: Competence Center of Food and Fermentation Technologies
Projects: HOST-PAR
Institutions: INSTITUTO FEDERAL FARROUPILHA
Projects: COVID-19 Disease Map
Institutions: BIOASTER
Projects: COVID-19 Disease Map
Institutions: Fundacion, Progreso y Salud
Institutions: Latvia University of Agriculture
Dr. Noelia Pérez-Rodríguez holds a PhD in Agro-Food Science and Technology and has over 6 years of expertise in the areas of Biomass Conversion and Bioprocess Technology with focus on the development of biorefinery strategies for total and sustainable conversion of biomass (lignocellulosic materials) into biobased products. Dr. Pérez-Rodríguez’s research interests include: i) biomass pretreatment and hydrolysis using chemical, physico-chemical or biological methods; ii) biomass conversion by ...
Projects: CC-TOP
Institutions: Technische Universität Darmstadt
PhD in Biotechnology, Research Associate at Department of Biotechnology and Systems Biology, National Institute of Biology
Projects: Rhodolive
Institutions: University of Kassel
Projects: INDIE - Biotechnological production of sustainable indole
Institutions: UNIBI: Bielefeld University
Projects: TRANSLUCENT
Institutions: Institute of Physiology Academy of Sciences
Translucent
Projects: Data science Übung
Institutions: Technical University of Munich
Projects: CoolWine
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-3485-1634
https://orcid.org/0000-0002-3485-1634
PhD candidate working on the CoolWine project
Projects: EmPowerPutida
Institutions: BASF
I am a research scientist in the Microbiology group at BASF. My expertise is the microbial production of secondary metabolites. I have more than 5 years of experience in engineering Actinomycetes and Pseudomonas strains with a strong focus on biosynthetic cluster expression.
Projects: MESI-STRAT
Institutions: German Cancer Research Center (DKFZ)
Projects: COVID-19 Disease Map
Institutions: Integrative BioInformatics Inc.
 https://orcid.org/0000-0001-7979-2386
https://orcid.org/0000-0001-7979-2386
Location: Mountain View, CA, USA
My research interest is in studying cellular and molecular pathways of COVID-19 disease.
Projects: SulfoSys, SulfoSys - Biotec
Institutions: University of Sheffield
Projects: COVID-19 Disease Map
Institutions: Gladstone Institutes
 https://orcid.org/0000-0001-5706-2163
https://orcid.org/0000-0001-5706-2163
Bioinformatics core director at Gladstone Institutes. Executive director of the National Resource for Network Biology. Co-founder of WikiPathways. Co-developer of Cytoscape.
Projects: BioDynamics
Institutions: Universitat Politècnica de València
 https://orcid.org/0000-0003-4144-3521
https://orcid.org/0000-0003-4144-3521
Jesús Picó is Full Professor of Automatic Control at The Department of Systems Engineering and Control of the Universitat Politècnica de València (UPV). In 2013 he created the Laboratory of Synthetic Biology and Biosystems Control (SB2CLab) of the Institute of Automation and Industrial Informatics (ai2) at the UPV, the first multidisciplinary lab in Spain integrating systems and control engineers, bioinformatics and biotechnologists. His research interests are in the application of systems ...
Projects: MESI-STRAT
Institutions: De Duve Institute
Projects: TRALAMINOL
Institutions: Technische Universität Darmstadt
Project Manager
Projects: COVID-19 Disease Map
Institutions: Hospital del Mar Research Institute (IMIM)
 https://orcid.org/0000-0003-1244-7654
https://orcid.org/0000-0003-1244-7654
Postdoctoral Researcher
Projects: ComPASs - Common Pathways in Amyotrophic Lateral Sclerosis (ALS) and Spinal Muscular Atrophy (SMA), miRiAD - exploring the role of microRNAs in T cell function and anti-viral defence, LungCARD - Blood test for clinical therapy guidance of non-small cell lung cancer patients, Mapping Disease Modules Overlaps in Biological Networks
Institutions: RNA Systems Biology Lab - BioISI/FCUL
 https://orcid.org/0000-0002-4217-0054
https://orcid.org/0000-0002-4217-0054
Projects: PSYSMO
Institutions: Imperial College London
Projects: FAIRDOM user meeting
Institutions: CSIC
Projects: SysMetEx
Institutions: Università della Svizzera Italiana
Projects: AquaHealth (ERA-BlueBio)
Institutions: Department of Planning, Aalborg University
 https://orcid.org/0000-0002-7462-2668
https://orcid.org/0000-0002-7462-2668
Projects: Rhodolive, Establishing an innovative and transnational feed production approach for reduced climate impact of the aquaculture sector and future food supply, Establishing an innovative and transnational feed production approach for reduced climate impact of the aquaculture sector and future food supply
Institutions: Leuphana University of Lüneburg, Institut for Food and Environmental Research (ILU), Institute for Food and Environmental Research (ILU)
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Projects: BioTarget Database
Institutions: University of Ljubljana
 https://orcid.org/0000-0002-8429-0273
https://orcid.org/0000-0002-8429-0273
Projects: Kinetics on the move - Workshop 2016, COVID-19 Disease Map, NMTrypI - New Medicines for Trypanosomatidic Infections, CoVIDD - Coronavirus interactions in drug discovery - optimization and implementation
Institutions: Kinetics on the move Workshop at HITS, Heidelberg Institute for Theoretical Studies, University of Eastern Finland (UEF)
 https://orcid.org/0000-0002-2801-8902
https://orcid.org/0000-0002-2801-8902
Projects: CC-TOP
Institutions: University of Groningen
Projects: SysMetEx
Institutions: Ruhr University Bochum
Projects: COVID-19 Disease Map
Institutions: Barcelona Supercomputing Center
 https://orcid.org/0000-0002-7496-844X
https://orcid.org/0000-0002-7496-844X
Projects: SUMO
Institutions: University of Sheffield
Work in my laboratory is focussed on microbial physiology - the study of how bacteria and other microorganisms work. Although rooted in the tradition of bacterial growth and intermediary metabolism, microbial physiology now embraces molecular biology, genetics, biochemistry, and indeed any discipline that can shed light on bacterial function. Much of our experimental work is conducted with Escherichia coli, the pre-eminent ‘model’ organism with unrivalled ease of genetic and physiological ...
Projects: KOSMOBAC
Institutions: University of Groningen
Bert Poolman is professor in biochemistry and program director of the Centre for Synthetic Biology. His research focuses on the elucidation of the mechanisms by which signals are transduced and small molecules are translocated across cellular membranes. Cells are reengineered for the production of correctly folded membrane proteins, and methods are developed to reconstitute complex molecular assemblies in synthetic membranes and to analyze their functional and structural properties. The in vitro ...
Projects: BESTER
Institutions: Green Biologics Ltd
Projects: COVID-19 Disease Map
Institutions: The European Bioinformatics Institute < EMBL-EBI
Projects: MetApp
Institutions: INSA Toulouse
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: SysVirDrug
Institutions: University of Leiden
Projects: WG2 - Integration of early phase studies and low environmental impact actions, WG6 - Promote the transfer of knowledge, WG5 - Promote dissemination, WG1 - Compound libraries coordination and integration of compound design, BioTarget Database, Compound Data
Institutions: Università degli Studi di Siena
 https://orcid.org/0000-0003-2574-3911
https://orcid.org/0000-0003-2574-3911
Projects: Hi-IMPAcTB, MIT SRP
Institutions: Massachusetts Institute of Technology
Researcher at BioMIcro Center, Data Management and Analysis Core; Massachusetts Institute of Technology
Projects: Ultreia
Institutions: Universidade de Santiago de Compostela
Projects: MESI-STRAT
Institutions: German Cancer Research Center (DKFZ)
 https://orcid.org/0000-0002-9659-9825
https://orcid.org/0000-0002-9659-9825
Projects: SEEK tutorial for DigiSal, DigiSal
Institutions: Norwegian University of Life Sciences
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
PSYSMO project manager
Projects: COVID-19 Disease Map
Institutions: ICESI University
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Julio is the leader of the Computational Biology Lab at MBG-CSIC (Pontevedra, Spain), an institute of CSIC (Spanish National Research Council). More info at https://www.bangalab.org
Our research is focused in computational systems and synthetic biology.
We use mathematical modelling, simulation and optimization to understand complex biological systems and processes.
Current research topics include:
-
Reverse engineering: systems identification (dynamic modelling) of biological networks
...
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur de Tunis
Projects: Working Group Nicole Radde, SteaPKMod
Institutions: University of Stuttgart
 https://orcid.org/0000-0002-5145-0058
https://orcid.org/0000-0002-5145-0058
Projects: COVID-19 Disease Map
Institutions: University of Montpellier
 https://orcid.org/0000-0001-6453-5707
https://orcid.org/0000-0001-6453-5707
Leader of Computational Systems Biology team at LPHI UMR 5235 CNRS and University of Montpellier
Projects: COSMIC
Institutions: Wageningen University & Research
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Duesseldorf
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: National University of Ireland
Projects: COVID-19 Disease Map
Institutions: Mansoura Faculty of Medicine
Projects: COVID-19 Disease Map
Institutions: University of Bonn
Projects: TRANSLUCENT
Institutions: University of Cordoba
Professor of Microbiology
Projects: PSYSMO
Institutions: CSIC Granada
Projects: STREAM
Institutions: University of Warwick
Projects: DigiSal-BT8121
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-0754-0507
https://orcid.org/0000-0002-0754-0507
Projects: Dynamic interplay between nuclei and mitochondria in ageing cells
Institutions: University of Copenhagen
Projects: KOSMOBAC
Institutions: University of Aberdeen
I am postdoc in the group of Prof Ian Booth in Aberdeen working on the biochemical and biophysical characterisation of bacterial channels.
Projects: Auromega
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-0607-2561
https://orcid.org/0000-0002-0607-2561
Projects: SysVirDrug
Institutions: Technology Transfer Heidelberg GmbH
Projects: MESI-STRAT
Institutions: University of Groningen
Projects: COVID-19 Disease Map
Institutions: INSERM
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: ODDITY, VIALACTEA, FATCRACK
Institutions: Universidade de Santiago de Compostela
 https://orcid.org/0000-0001-8227-6665
https://orcid.org/0000-0001-8227-6665
Projects: HubMOL
Institutions: Josep Carreras Leukaemia Research Institute (IJC)
 https://orcid.org/0009-0005-7220-690X
https://orcid.org/0009-0005-7220-690X
Projects: Group Microbial Interface Biology
Institutions: Forschungszentrum Borstel
Projects: FAIRDOM user meeting
Institutions: Imperial College London
Projects: MIX-UP
Institutions: Tsinghua University
Projects: NIRBTEST
Institutions: Institut Curie
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0003-3851-9978
https://orcid.org/0000-0003-3851-9978
Projects: PSYSMO, MOSES, COSMIC, BaCell-SysMO
Institutions: University of Stuttgart
Professor for Biochemcial Engineering, University Stuttgart
Projects: STREAM
Institutions: University of Tuebingen
Projects: MESI-STRAT Review
Institutions: University of Innsbruck
 https://orcid.org/0000-0003-3540-0402
https://orcid.org/0000-0003-3540-0402
Projects: MESI-Review 2024
Institutions: University of Innsbruck
Reviewer for MESI-STRAT project 2024
Projects: SysMO-LAB, de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, FAIRDOM, EnzymeML, FAIRDOM Community Workers, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", MIX-UP, COVID-19 Disease Map, ERNEST Mapping Group Pilot Study, CC-TOP, SABIO-VIS, SDBV/HITS
Institutions: Heidelberg Institute for Theoretical Studies
Within the de.NBI project my functions in the de.NBI-SysBio node comprise content curation, requirements elicitation, and community engagement for the users of biochemical reaction kinetics database SABIO-RK as well as of the data management platform SEEK.
Projects: Cassava Gates Curation
Institutions: General Bioinformatics Ltd
Projects: COVID-19 Disease Map
Institutions: Fundación Progreso y Salud
 https://orcid.org/0000-0003-4314-7208
https://orcid.org/0000-0003-4314-7208
Projects: KOSMOBAC
Institutions: University of Aberdeen
My background is physics engineering & biomedical engineering. I did my PhD in Surrey on the modelling of response of mammalian cells to radiation of different qualities. I have been working at the University of Aberdeen since November 2007 as a theoreticien research fellow of the KOSMOBAC project. We are investigating the homeostasis of ions in bacteria E. coli. I have been working at a model of the buffering capacity of the cytopplasm, arising from the presence of weak acids and bases. We ...
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: COSMIC
Institutions: University of Ulm
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: FAIRDOM
Institutions: ETH Zurich & Basel / The Scientific IT Services (SIS) division
Head of Scientific IT Services and member of the ITS executive board at ETH Zurich. Project manager of SyBIT. Project partner of the FAIRDOM Initiative.
Projects: COVID-19 Disease Map
Institutions: Gladstone Institutes
Projects: CC-TOP
Institutions: Johnson Matthey
Department of Microbiology, Faculty of Pharmacy. University of Granada (Spain).
Projects: COVID-19 Disease Map
Institutions: IBM Research, Zurich
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Innsbruck
I am a postdoctoral researcher in the group of Julio Banga. My research is focused on computational systems biology with particular attention to the mathematical modelling of biosystems and bioprocesses. Some of the topics we address are:
- Parameter estimation
- Model identifiability
- Global sensitivity analysis
- Optimal experimental design
- Dynamic optimization
- Robust control of diffusion-reaction systems
Projects: STREAM
Institutions: INBIOTEC Leon
Projects: PSYSMO
Institutions: CSIC Madrid
Projects: SilicoTryp
Institutions: University of Edinburgh
I am a Postdoc at Keith Matthews lab in the Institute of Immunology and Infection Research, Edinburgh University. As part of the SilicoTryp project we are in charge of performing Targeted disruption and Overexpression of critical enzymes of Trypanosoma brucei redox metabolism enzymes and developmental perturbations to provide part of the necessary data for the construction of the model. Also generate consistent samples, so that data can be integrated and quantification results are guarateed to ...
Institutions: Heidelberg Institute for Theoretical Studies
Projects: SUMO
Institutions: University of Sheffield
I am a post-doctoral research associate working in Sheffield in the SUMO consortium. My research focuses on transcriptional regulation in E. coli, with particular emaphasis on the transcriptomic analysis of steady-state chemostat cultures using both microarray and qRT-PCR approaches.
Previous experience, especially that gained during my PhD, involved work on Salmonella physiology and lag phase growth, focusing particularly on gene-expression and transcriptional regulation. Other techniques used ...
Projects: BESTER
Institutions: PROCESSIUM
Process engineer since 2011 at Processium
Projects: KOSMOBAC
Institutions: University of Aberdeen
physicist
Projects: COVID-19 Disease Map
Institutions: Institut Pasteur de Tunis
Projects: PSYSMO
Institutions: University of Dortmund
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
Projects: DigiSal, GenoSysFat, DigiSal-BT8121
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-1702-7379
https://orcid.org/0000-0002-1702-7379
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: Heinrich Heine University of Düsseldorf
Projects: GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
Projects: Sustainable co-production
Institutions: Indear
Projects: SysMO Funders
Institutions: Forschungszentrum Jülich
Working for Project Management Jülich, I am responsible for the SysMO-Office. This implies the coordination of communication between funding organisations (i.e. the Steering Committee), the Scientific Advisory Board, the Data Management Group, the PALs and the research groups.
Additionally, I am one of the administrative contacts for all German groups at Project Management Jülich.
Before I came to Jülich, my research focus was modelling and simulation in marine ecosystems.
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
Projects: SEEK tutorial for DigiSal, DigiSal
Institutions: Norwegian University of Life Sciences
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München / Institute of Experimental Genetics
Projects: SAFE-Aqua
Institutions: National Center for Genetic Engineering and Biotechnology
Projects: From waste to taste: exploring innovative food applications of postharvest fish losses_WASTE2TASTE, BlueTekna research group
Institutions: Department of Ecosustainable Marine Biotechnology, Stazione Zoologica Anton Dohrn, Department of Ecosustainable Marine Biotechnology, Department of Ecosustainable Marine Biotechnology, Stazione Zoologica Anton Dohrn
Institutions: University of Stavanger
Physical chemist with expertise in experimental kinetics, molecular biology, and mathematical modeling.
Projects: COVID-19 Disease Map
Institutions: University of Rome Tor Vergata
Projects: COVID-19 Disease Map
Institutions: University of Heidelberg
 https://orcid.org/0000-0002-8552-8976
https://orcid.org/0000-0002-8552-8976
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: University Hospital Heidelberg
Projects: Kinetics on the move - Workshop 2016, de.NBI-SysBio
Institutions: University of Heidelberg
Ph.D. student at the Norwegian University of Life Sciences interested in protein stoichiometry and genetics. Currently, I am studying nicotinic acetylcholine receptors in salmon lice, in an effort to understand how they function, their stoichimetric structure and their importance to the salmon louse. This is part of a larger fundamental research project which covers several invertebrates including bees, mosquitos and ticks, as ...
Projects: PhotoBoost
Institutions: Instituto de Tecnologia Química e Biológia - Universidade Nova de Lisboa
Projects: FAIRDOM user meeting
Institutions: Nazarbayev University
Projects: Sustainable co-production
Institutions: BIOCERES S.A.
Projects: PSYSMO
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: SynBio4Flav, PROMISEANG
Institutions: CSIC, Spanish National Center for Biotechnology (CNB-CSIC)
 https://orcid.org/0000-0001-8138-500X
https://orcid.org/0000-0001-8138-500X
Projects: FAIRDOM
Institutions: Heidelberg Institute for Theoretical Studies
Projects: COVID-19 Disease Map
Institutions: Autonomous University of Madrid
Projects: Xenophiles Systems Biology
Institutions: VU University Amsterdam
Committee member Ph D thesis Joost Aerts
Projects: SysMetEx
Institutions: Universitity Duisburg-Essen
Projects: COVID-19 Disease Map
Institutions: Harvard Medical School
Projects: Sustainable co-production
Institutions: Wageningen University & Research
Projects: _p_stRT
Institutions: Indiana University Bloomington
Projects: SUMO
Institutions: University of Edinburgh
Projects: CF transcriptome
Institutions: RNA Systems Biology Lab - BioISI/FCUL
Projects: BaCell-SysMO
Institutions: University of Greifswald
I am PhD student at Prof.Uwe Voelker lab in Department of Functional Genomics. My area of research is microbial functional genomics in particular analysing the whole transcriptome(by microarray and other molecular biolology methods) of B.subtilis under various stress conditions. I use QconCAT strategy for absolute quantification of carbon metabolic enzymes via MRM(multiple reaction monitoring) by LC-MS/MS. I also perofrm experiments for understanding of dynamics of SigmaB network for modelling.
Projects: COVID-19 Disease Map
Institutions: Guy's and St Thomas' NHS Foundation Trust and King's College London
Projects: Environmentally-friendly bioadhesives from renewable resources (WooBAdh)
Institutions: University of Ljubljana
Projects: FAIRDOM user meeting
Institutions: JAPAN TOBACCO INC
Dr. Venkata Satagopam is a Group Leader, Senior Research Scientist and Deputy Head of Bioinformatics core facility, LCSB, University of Luxembourg; Technical Coordinator (TeC) of ELIXIR-Luxembourg Node and CTO & Co-founder ITTM S.A. Luxembourg. Before joining LCSB, between 2004-2012 he worked as a Senior Bioinformatics Scientist at the EMBL, Heidelberg. Before EMBL he worked as a Bioinformatics Scientist at LION bioscience AG, Heidelberg. He is having more than 20 years of working experience ...
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: SysMilk
Institutions: ETH Zurich
Projects: SUMO
Institutions: University of Stuttgart
From 2005 to 2008 I was group leader at the Institute for System Dynamics at the University of Stuttgart. Since 2008 I am now Professor of Systems Biology at the University of Luxembourg.
The research of the Systems Biology Group at the University of Luxembourg is focussed in the area of experimental and theoretical systems biology. We are applying different modelling techniques (mainly ODE and logical) to biological systems to develop suitable computational models. The analysis of these models ...
Projects: SUMO
Institutions: University of Stuttgart
PI of the SUMO work package "Detailed kinetic modelling, identification and analysis of the citric acid cycle and respiratory chains of E. coli". Director of the Institute for System Dynamics, Universität Stuttgart, Germany.
Projects: OXYMOD
Institutions: Norwegian University of Science and Technology
My group investigates dynamic regulation and control mechanisms of cellular signal transduction networks by a combination of theoretical, experimental and computational methods. We seek to make sense of our biological data with the help of mathematical models, which ideally enable us to make valid predictions for new experiments, thereby generating novel biological insights.
Projects: Cell4Chem
Institutions: Helmholtz Zentrum für Umweltforschung (UFZ)
Projects: COVID-19 Disease Map
Institutions: esqLABS GmbH
Projects: COVID-19 Disease Map, Molecular characterization and angiogenic properties of extracellular vesicles in healthy and preeclamptic amniotic fluid, Overexpression of microRNAs miR-25-3p, miR-185-5p and miR-132-3p in Late Onset Fetal Growth Restriction, Validation of Results and Study of the Biochemical Pathways Involved, Comparative Proteome Profile of Placental Sub-anatomical regions, The deregulated interplay of complement system and antioxidant activity as a driver of adverse pregnancy outcomes
Institutions: University of Rostock, Instituto de Investigación Sanitaria La Fe
 https://orcid.org/0000-0002-0034-7755
https://orcid.org/0000-0002-0034-7755
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: ETH Zurich
Projects: IMOMESIC
Institutions: Universitätsklinikum Leipzig
Projects: COSMIC
Institutions: University of Ulm
Dr. Bettina Schiel-Bengelsdorf Senior Scientist Department of Microbiology and Biotechnology University of Ulm 89069 Ulm, Germany
Projects: Hi-IMPAcTB, MIT SRP
Institutions: Massachusetts Institute of Technology
Projects: SulfoSys
Institutions: University of Vienna
Geneticist and Microbiologist with research focus on Archaea. Professor of Genetics in Ecology (Faculty of Life Sciences/University of Vienna, Austria) and associate professor of the Center of Geobiology (University of Bergen/Norway).
Projects: SysVirDrug
Institutions: ETH Zurich
Projects: PSYSMO
Institutions: University of Dortmund
Projects: COVID-19 Disease Map
Institutions: University Maastricht
Carsten Oliver Schmidt is Deputy head of the Department of SHIP/ Clinical-Epidemiological Research. He leads the functional division “Quality in the health Sciences”. The investigator is PI of several methods and research quality related projects, and speaker of the methods section of the German Society for Epidemiology. He works in the field of research methods, quality management, musculosceletal disorders, among others.
Projects: SCaRAB
Institutions: University of Greifswald
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: VU University Amsterdam
I am a beginning PhD student at the VU in Amsterdam and study the heterogeneity of yeast cells at near zero growth conditions. I have a versatile background in Biophysics and Systems Biology.
Detlef Schmiedl (PhD) is working at Fraunhofer ICT since 02/2006 as scientist & is project leader as well as senior scientist at Fraunhofer ICT. He is a chemist & has long term experience in processing of lignocellulose & chemical modification/ tailored functionalization of tannin & lignin as well as in base catalyzed chemical depolymerization of several lignin types into monomer & oligomer oxy-aromatic building blocks. Additionally he has long term experiences in analytical ...
Projects: COVID-19 Disease Map
Institutions: Helmholtz Zentrum München
Projects: Not specified
Institutions: Not specified
Creative Diagnostics is a leading manufacturer and supplier of antibodies, viral antigens, innovative diagnostic components, and critical assay reagents. In addition to providing contract R&D and biologic manufacturing services for diagnostic manufacturers along with GMP biologics manufacturing for the biopharmaceutical market, the company aims to continue to act as a trusted source for all researchers’ assay development and manufacturing needs.
Projects: PSYSMO
Institutions: University of Braunschweig
Projects: Alters-Krebs-Korrelation anhand der Patienteneingänge 2024
Institutions: BIDS
Projects: Core Facility Fluorescence Cytometry
Institutions: Forschungszentrum Borstel
Projects: SulfoSys, PSYSMO, SulfoSys - Biotec, EbN1 Systems Biology
Institutions: University of Braunschweig
Projects: SulfoSys - Biotec
Institutions: University of Braunschweig
Projects: Reduce Complexity (RCO) reconstruction
Institutions: Heinrich Heine University of Düsseldorf
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: Isolieren neuer Gluconobacter Stämme, 16S rRNA-Sequencing
Institutions: Technical University of Munich
Projects: MESI-STRAT
Institutions: University of Heidelberg
Projects: COVID-19 Disease Map
Institutions: Universität Konstanz
Projects: PSYSMO, SulfoSys, SulfoSys - Biotec
Institutions: University of Braunschweig
Projects: TRANSLUCENT
Institutions: University of Bonn
Projects: EmPowerPutida
Institutions: LifeGlimmer GmbH
Projects: C1Pro
Institutions: UNIBI: Bielefeld University
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
Projects: ZucAt
Institutions: University of Hohenheim
Projects: COSMIC
Institutions: Wageningen University & Research
PostDoc at Wageningen University, Laboratory of Microbiology
Projects: TRANSLUCENT
Institutions: University of Vienna
Projects: COVID-19 Disease Map
Institutions: Systems Biomedicine Center, University of Luxembourg
Projects: Not specified
Institutions: Not specified
With years of experience in the life sciences, Amerigo Scientific has integrated global cutting-edge technology resources to provide the latest technological equipment, instruments, biological products, and custom services for research institutions and enterprises in the biological, chemical, pharmaceutical, medical, and other industries.
Projects: COVID-19 Disease Map
Institutions: National Institute of Standards and Technology
Projects: Institut Pasteur's projects
Institutions: Institut Pasteur
 https://orcid.org/0000-0002-6301-7147
https://orcid.org/0000-0002-6301-7147
Projects: Clostridium beijerinckii pan-genome
Institutions: Brno University of Technology
 https://orcid.org/0000-0002-8269-4020
https://orcid.org/0000-0002-8269-4020
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: COMBINE Multicellular Modelling
Institutions: University of Florida
 https://orcid.org/0000-0002-4274-656X
https://orcid.org/0000-0002-4274-656X
Projects: BaCell-SysMO
Institutions: University of Erlangen-Nuernberg
I studied chemistry in Erlangen and started research on carbon catabolite regulation in Bacillus megaterium in my diploma thesis. During my PhD thesis I was concerned with quantitative analyses of protein-protein and protein-DNA interactions of CcpA, in vivo- and in vitro-characterization of point mutants of CcpA and coeffectors and I contributed to the structural analysis of different CcpA complexes. During a postdoctoral appointment I focused on interaction analyses of PrfA, a regulator of ...
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-6758-2979
https://orcid.org/0000-0002-6758-2979
Projects: COVID-19 Disease Map
Institutions: ICAHN Icahn School of Medicine at Mount Sinai
 https://orcid.org/0000-0002-1605-8130
https://orcid.org/0000-0002-1605-8130
Experienced in developing rule-based models of complex biochemical reaction networks, as well as developing new rule-based approaches. Extensive experience with BioNetGen, a tool for building complex reaction networks succinctly with rules. Currently working with Jonathan Karr at Dept. Genetics & Genomics at Mount Sinai School of Medicine on extending rule-based approaches to transcription, translation and other sequence-based mechanisms.
Projects: ERNEST Mapping Group Pilot Study
Institutions: Universitat Pompeu Fabra
Projects: SYLOBIO
Institutions: Rostock University Medical Centre
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
Projects: MESI-STRAT
Institutions: Vall d´Hebron Institute of Oncology (VHIO)
Luis Serrano did his PhD at the CBM (Madrid, Spain) on Cell Biology. Then he spent 4 years in the laboratory of Prof. A.R. Fehrs (MRC, UK) working in protein folding. In 1993, he became Group Leader at the EMBL (Heidelberg, Germany) working in Protein Folding and design. Ten years later, he was appointed head of the Structural & Computational Biology programme at the EMBL and he started to work on Systems Biology. By the end of 2006 he moved back to Spain to lead a programme working on Systems ...
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: COVID-19 Disease Map
Institutions: Indraprastha Institute of Information Technology Delhi
Projects: BESTER
Institutions: Imperial College London
Projects: COVID-19 Disease Map
Institutions: The European Bioinformatics Institute < EMBL-EBI
Projects: Noisy-Strep
Institutions: University of Groningen
Projects: COVID-19 Disease Map
Institutions: NYU Grossman School of Medicine
Projects: MESI-STRAT, Systems modelling age-related changes in the maintenance of dermal extra-cellular matrix: mechanisms and interventions, Are microRNAs key mediators of cartilage destruction in osteoarthritis?, Dynamic interplay between nuclei and mitochondria in ageing cells, Outreach - Breast Cancer RStudio-Shiny application, Outreach - Simulation of breast cancer development using an agent based modelling approach, Outreach - Senescence RStudio-Shiny application, Outreach - Simulation of cellular senescence using an agent based modelling approach, Selective Destruction in Ageing
Institutions: University of Newcastle
 https://orcid.org/0000-0003-3096-6386
https://orcid.org/0000-0003-3096-6386
Projects: Lipofungi, Bio4Fuels, LIGNOLIPP
Institutions: Norwegian University of Life Sciences
Projects: MOSES, PSYSMO, SysMO-LAB, SulfoSys
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: SUMO
Institutions: University of Amsterdam
Projects: Service to URV Tarragona, Spain with respect to their Safety Assessment of Endocrine Disrupting Chemicals model (Active NOW), EraCoBiotech 2 nd call proposal preparation, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation
Institutions: VU University Amsterdam, Universitat Rovira i Virgili
 https://orcid.org/0000-0001-9103-9127
https://orcid.org/0000-0001-9103-9127
Projects: MESI-STRAT
Institutions: The Arctic University of Norway (UiT)
Projects: TRALAMINOL
Institutions: Technische Universität Darmstadt
 https://orcid.org/0000-0002-0297-2191
https://orcid.org/0000-0002-0297-2191
Rahuman Sheriff is a Senior Project Leader (BioModels) at the European Bioinformatics Institute, European Molecular Biology Laboratory (EMBL-EBI), Hinxton, Cambridge, UK. He interests include mathematical modelling, development of novel tools and resources for building models, immune digital twin, quantitative imaging, single cell systems biology and chemoinformatics. He is one of the editors of Systems Biology Markup Language (SBML).
Projects: COVID-19 Disease Map
Institutions: Golestan University of Medical Sciences
 https://orcid.org/0000-0002-0773-1610
https://orcid.org/0000-0002-0773-1610
I would like to collaborate on the COVID-19 projects. I am currently a faculty member of the Infectious Diseases research center, at the Golestan University of Medical Sciences, Iran, and studying clinical information of patients with COVID-19. I am a microbiologist and can help other teams to evaluate the data obtained from COVID-19 cases.
Projects: SulfoSys, SulfoSys - Biotec, HOTSOLUTE, Computational pathway design for biotechnological applications, SCyCode The Autotrophy-Heterotrophy Switch in Cyanobacteria: Coherent Decision-Making at Multiple Regulatory Layers
Institutions: University Duisburg-Essen, Universitity Duisburg-Essen
 https://orcid.org/0000-0002-9905-541X
https://orcid.org/0000-0002-9905-541X
Head of the group of Molecular Enzyme Technology and Biochemistry (Faculty of Chemistry) at the University of Duisburg-Essen. My research interest is on archaeal physiology with a special focuss on the central carbohydrate metabolism of (hyper)thermophilic Archaea and its regulation. The aim is to gain a systems level understanding by the combination of modern highthrouput analyses with classical biochemistry and molecular biology. Archaea possess many novel enzymes and pathways and our aim is ...
Projects: SysMO-LAB
Institutions: University of Rostock
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: PSYSMO
Institutions: University of Stuttgart
Projects: SulfoSys
Institutions: Wageningen University & Research
Projects: BioZEment 2.0
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0001-9413-1623
https://orcid.org/0000-0001-9413-1623
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-4989-5311
https://orcid.org/0000-0003-4989-5311
Projects: Auromega
Institutions: Norwegian University of Science and Technology
Projects: Sustainable co-production
Institutions: Indear
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Leiden
Gopakumar Sivasankarapillai (Ph.D.) is a member of the research group of the chair of Forest Biomaterials, Freiburg material research center, the University of Freiburg since 2014. I am involving, various routes for the synthesis of lignin and tannin derivatives, various chemical modifications of cellulose and xylans for the design of functional composite materials.
Projects: Rhodolive
Institutions: National Institute of Chemistry (Slovenia)
Projects: TRALAMINOL
Institutions: Technische Universität Darmstadt
 https://orcid.org/0000-0002-2150-350X
https://orcid.org/0000-0002-2150-350X
Postdoctoral fellow
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0002-5901-1404
https://orcid.org/0000-0002-5901-1404
I am a multicell and multiscale modeler and member of the Biocomplexity group at Indiana University. The group produces and maintains the multicellular modelling package Compucell3D. I am also a voting member of the USA committee to the International Organization for Standards (ISO) working group on standards in biotechnology and the chairman of a sub-sub group on publishing standards.
Projects: Xenophiles Systems Biology
Institutions: Wageningen University & Research
Professor at WUR Member of Doctorate Committee Joost Aerts
Projects: BESTER
Institutions: Green Biologics Ltd
Projects: STREAM
Institutions: University of Aberdeen
Projects: COVID-19 Disease Map
Institutions: Luxembourg Centre for Systems Biomedicine (LCSB)
Projects: SysVirDrug
Institutions: Leiden University Medical Center
Projects: DigiSal, GenoSysFat
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-2883-861X
https://orcid.org/0000-0002-2883-861X
Projects: PSYSMO, MOSES, SysMO DB, SysMO-LAB, SulfoSys, SulfoSys - Biotec, Whole body modelling of glucose metabolism in malaria patients, FAIRDOM, Molecular Systems Biology, COMBINE Multicellular Modelling, HOTSOLUTE, Steroid biosynthesis, Yeast glycolytic oscillations, Computational pathway design for biotechnological applications, SCyCode The Autotrophy-Heterotrophy Switch in Cyanobacteria: Coherent Decision-Making at Multiple Regulatory Layers, Project Coordination, WP 3: Drug release kinetics study, Glucose metabolism in cancer cell lines
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester, University of Stellenbosch, University of Manchester - Department of Computer Science, Stellenbosch University
Projects: MIT SRP, Hi-IMPAcTB, MetNet
Institutions: Massachusetts Institute of Technology
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
Projects: SysMO-LAB
Institutions: Norwegian University of Life Sciences
Projects: COVID-19 Disease Map
Institutions: Oregon Health & Science University (OHSU)
Projects: MESI-STRAT
Institutions: PATH - Patients' Tumor Bank of Hope
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
Projects: FAIRDOM user meeting
Institutions: Standigm
Projects: Noisy-Strep
Institutions: University of Groningen
Projects: MetApp
Institutions: Sinvent AS - SINTEF
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Freiburg
Projects: CropClock
Institutions: Syngenta Seeds GmbH
Projects: iRhythmics
Institutions: Rostock University Medical Centre
Projects: SUMO
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: SysMO DB, Whole body modelling of glucose metabolism in malaria patients, Manchester Institute for Biotechnology, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, GenoSysFat, DigiSal, FAIRDOM user meeting
Institutions: University of Manchester - Department of Computer Science, Manchester Centre for Integrative Systems Biology, University of Manchester
 https://orcid.org/0000-0003-4958-0184
https://orcid.org/0000-0003-4958-0184
Interested in systems + synthetic biology, biotechnology, mountaineering, swimming, running, and the occasional cup of tea. Once diagnosed as an ENFP.
Projects: SulfoSys - Biotec
Institutions: University of Braunschweig
Projects: COMBINE Multicellular Modelling
Institutions: Technische Universität Dresden
 https://orcid.org/0000-0003-3649-2433
https://orcid.org/0000-0003-3649-2433
Projects: RNA-seq data for paper, RNA-seq data
Institutions: Previwo AS, Norwegian University of Life Sciences
Projects: Kinetics on the move - Workshop 2016, COVID-19 Disease Map
Institutions: Kinetics on the move Workshop at HITS, University of Edinburgh
Projects: SYSTERACT
Institutions: University of Tübingen
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: University of Freiburg
Projects: BaCell-SysMO
Institutions: University of Greifswald
I started to work with B. subtilis during my diploma thesis in Marburg, analyzing the gene expression pattern during sporulation and their control by the four sporulation sigma factors. This work was continued during my PhD thesis in Greifswald. In collaboration with Prof. Bremer and Prof. Marahiel in Marburg we also studied additional adaptation processes of B. subtilis, like the adaptation to low temperatur and high osmolarity. I am now working as a staff scientist in Prof. Völkers lab in ...
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: BaCell-SysMO
Institutions: University of Erlangen-Nuernberg
Projects: SUMO
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: IMOMESIC
Institutions: German Cancer Research Center (DKFZ)
Projects: COVID-19 Disease Map
Institutions: Ontario Institute for Cancer Research
 https://orcid.org/0000-0002-4650-631X
https://orcid.org/0000-0002-4650-631X
Projects: OXYMOD
Institutions: Norwegian University of Life Sciences
Projects: Stress granules
Institutions: University of Bergen
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: HOTSOLUTE
Institutions: University Duisburg-Essen
Projects: BaCell-SysMO
Institutions: University of Newcastle
Projects: GenoSysFat, SEEK tutorial for DigiSal, DigiSal
Institutions: Norwegian University of Life Sciences
Projects: FAIRDOM
Institutions: ETH Zurich & Basel / The Scientific IT Services (SIS) division
Mag. Annette Strauch-Davey, M-A.
Annette Strauch-Davey (M.A.) is a cultural scientist / European ethnologist and English literature and medieval studies scholar from the Georg-August University of Göttingen. She had lived in Cardiff and Aberystwyth in Wales for 15 years after her studies (including work at the Museum of Welsh Life and the National Library of Wales). For more than 10 years she has been interested in and dealing with the research data of scientists of various disciplines, also as ...
Projects: Cell4Chem
Institutions: Jožef Stefan Institute
Projects: Bergen(Ziegler lab) project AF-NADase
Institutions: University of Bergen
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: BaCell-SysMO
Institutions: University of Goettingen
Associate Professor at Wageningen University & Research
Researcher at the University Medical Center Göttingen, Department of Medical Informatics
Projects: INBioPharm, SYSTERACT, BESTER, OXYMOD
Institutions: SINTEF
Projects: COVID-19 Disease Map
Institutions: Ankara University
 https://orcid.org/0000-0001-6274-3385
https://orcid.org/0000-0001-6274-3385
Projects: COVID-19 Disease Map
Institutions: Imperial College London
Projects: SCaRAB
Institutions: Norwegian University of Science and Technology
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-1870-0104
https://orcid.org/0000-0003-1870-0104
Projects: EnzymeML
Institutions: University of Manchester - Department of Computer Science
 https://orcid.org/0000-0001-7020-1236
https://orcid.org/0000-0001-7020-1236
Projects: STREAM
Institutions: University of Groningen
Projects: TRANSLUCENT
Institutions: Institute of Physiology Academy of Sciences
biochemistry, molecular biology and physiology of lower eukaryotes, characterization of cell membrane transporters
Projects: PlaSMo model repository
Institutions: University of Edinburgh
Projects: TRALAMINOL
Institutions: Technische Universität Darmstadt
Projects: Lipofungi, Bio4Fuels, DeepHyperSpec, LIGNOLIPP
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-6015-0893
https://orcid.org/0000-0001-6015-0893
Projects: Mass spectrometry proteomics for biomarker discovery, Supplementary Information 2 associated with the manuscript entitled " Label free Mass spectrometry proteomics reveals different pathways modulated in THP-1 cells infected with therapeutic failure and drug resistance Leishmania infantum clinical isolates", Supplementary Information 2 associated with the manuscript entitled "Label free Mass spectrometry proteomics reveals different pathways modulated in THP-1 cells infected with therapeutic failure and drug resistance Leishmania infantum clinical isolates", MS identification of L infantum proteins related to their drug resistance patterns for new drug targets identification and ecotoxicological evaluations of their environmental and interspecies impact, Enhanced Anticancer Effect of Thymidylate Synthase Dimer Disrupters Promoting Intracellular Accumulation, Biochemical characterization of the feedforward loop between CDK1 and FOXM1 in epidermal stem cells, Drug Discovery and Biotechnology Standard Operating Procedures, Defining the role of fibroblasts in skin expansion, Proteo-Lipidomics Serum Signatures associated with Bevacizumab response in High Grade Serous Ovarian Carcinoma
Institutions: Università degli Studi di Modena e Reggio Emilia (UNIMORE), Università di Modena e Reggio Emilia, Life Sciences, Life Sciences, Università di Modena e Reggio Emilia
 https://orcid.org/0000-0002-7244-7106
https://orcid.org/0000-0002-7244-7106
Junior researcher in Medicinal Chemistry
Projects: Systems toxicology of Atlantic cod
Institutions: University of Oslo
Projects: Rhodolive
Institutions: Düzen Biological Sciences R&D and Production Company
Projects: STREAM
Institutions: University of Groningen
Projects: BioRECIPE representation format
Institutions: University of Pittsburgh
Projects: EmPowerPutida
Institutions: CSIC
CSIC Madrid, CSIC
Projects: COVID-19 Disease Map
Institutions: Swiss Institute of Bioinformatics
Projects: SYLOBIO
Institutions: Universitätsmedizin Greifswald
 https://orcid.org/0009-0006-5141-2154
https://orcid.org/0009-0006-5141-2154
Projects: PhotoBoost
Institutions: University of Lancaster
Projects: Not specified
Institutions: Not specified
BlueVision Technologies Europe GmbH (BVT) is a leading Germany based company, The company is sister concern of "BalaJi MicroTechnologies Pvt. Ltd. India". We are privately held & unit of "B.B. Group of Companies". The group is into business since 1949.
"We are supplier of high performance Machine Vision & Line Scan Cameras & precision Machine Vision lenses under our brand name "BMT" which is also known as "BMT Machine Vision lens ". Our company core interest lies in ITS/surveillance, ...
Projects: COVID-19 Disease Map
Institutions: KAUST
Institutions: University of Amsterdam
Professor Quantitative Microbial Physiology Dept Mol Mic Phys Swammerdam Inst Life Sc University of Amsterdam
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0002-4296-5068
https://orcid.org/0000-0002-4296-5068
Projects: MESI-STRAT
Institutions: PD-value B.V.
Projects: FAIRDOM user meeting
Institutions: Lund University
Projects: Xenophiles Systems Biology
Institutions: Utrecht University
PhD student @ "Quantitative Microbial Phenotyping" Institute of Bio- and Geosciences, IBG-1: Biotechnology Forschungszentrum Jülich GmbH 52425 Jülich, Germany
Projects: SUMO
Institutions: University of Amsterdam
Projects: SysMO-LAB, SysMilk, IMOMESIC
Institutions: Wageningen University & Research, VU University Amsterdam
 https://orcid.org/0000-0003-3929-0423
https://orcid.org/0000-0003-3929-0423
Since August 2008 I am professor in Systems Biology at the VU University Amsterdam. My Systems Bioinformatics group focusses on systems biology with a special focus on integrative bioinformatics. It aims at forming bridges between the classical bottom-up approaches in systems biology and the more data-driven approaches in classical bioinformatics. We combine experimental, modeling and theoretical approaches to study cellular physiology, with an emphasis on metabolic networks.
Projects: COVID-19 Disease Map
Institutions: Pacific Northwest National Laboratory
Projects: COVID-19 Disease Map
Institutions: University of Rochester
Prof. Dr. Kathrin Thedieck MESI-STRAT Coordinator Leader WP8 – Project Coordination
Projects: CC-TOP, TRALAMINOL
Institutions: Technische Universität Darmstadt
Projects: Not specified
Institutions: Not specified
Ace Therapeutics is a preclinical contract research provider dedicated to offering comprehensive one-stop services. Our expertise spans across various preclinical services, including efficacy testing, pharmacokinetics, toxicology, and biomarker development, to support clients in their research and development efforts.
Projects: KOSMOBAC
Institutions: University of Aberdeen
Associate Professor for Systems Biomedicine, Luxembourg Centre for Systems Biomedicine, University of Luxembourg
Projects: STREAM
Institutions: University of Aberdeen
Quantitative proteomics
Projects: GenoSysFat, DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0003-1415-4652
https://orcid.org/0000-0003-1415-4652
Projects: ImmPort - data sharing
Institutions: ImmPort
Projects: COSMIC
Institutions: University of Nottingham
Projects: BioZEment 2.0
Institutions: Oslo and Akershus University College
Projects: HubMOL
Institutions: UiT The Arctic University of Norway
 https://orcid.org/0009-0008-2364-5633
https://orcid.org/0009-0008-2364-5633
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Projects: COVID-19 Disease Map
Institutions: New York University
Projects: IMOMESIC
Institutions: German Cancer Research Center (DKFZ)
Projects: CC-TOP
Institutions: University of Goettingen
Projects: CoolWine
Institutions: Bodegas Roda S.A.
Projects: _p_stRT
Institutions: National Institute of Biology
Projects: CC-TOP
Institutions: University of London UCL
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: University of Freiburg
 https://orcid.org/0000-0003-2822-5191
https://orcid.org/0000-0003-2822-5191
Khasan Topiloff is an accomplished professional with a background in economics and IT. Since December 2022, he has been working as an IT specialist at the International Nordic University. Born on August 13, 1997, in Tashkent, Uzbekistan, Khasan completed his higher education in 2022 at Tashkent State University of Economics and Ural State University of Economics. Fluent in Russian and English, he has also held various positions in academia and the private sector, including roles at Tashkent State ...
Projects: MESI-STRAT
Institutions: PATH - Patients' Tumor Bank of Hope
Projects: FAIRDOM user meeting
Institutions: Vall d'Hebron Institut de Recerca (VHIR)
Projects: ExtremoPharm
Institutions: Immanuel Kant Baltic Federal University (IKBFU)
Projects: COVID-19 Disease Map
Institutions: Earlham Institute
 https://orcid.org/0000-0002-5600-8189
https://orcid.org/0000-0002-5600-8189
Projects: FAIRDOM user meeting
Institutions: Peter MacCallum Cancer Centre
Projects: Thrust C2
Institutions: Heidelberg Institute for Theoretical Studies
Projects: COVID-19 Disease Map
Institutions: Liverpool John Moores University
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Freiburg
Projects: PSYSMO
Institutions: University of Hannover
Projects: COVID-19 Disease Map
Institutions: European Molecular Biology Laboratory
 https://orcid.org/0000-0002-7249-9379
https://orcid.org/0000-0002-7249-9379
Projects: COVID-19 Disease Map
Institutions: Monash University
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: John Innes Centre
Projects: SAFE-Aqua
Institutions: National Center for Genetic Engineering and Biotechnology
 https://orcid.org/0000-0003-4710-2613
https://orcid.org/0000-0003-4710-2613
Projects: PSYSMO
Institutions: Helmholtz Centre for Infection Research Braunschweig
Projects: Millar group, PlaSMo model repository, PHYTOCAL: Phytochrome Control of Resource Allocation and Growth in Arabidopsis and in Brassicaceae crops, Light and plant development, Light control of leaf development, Toggle switch, Reduce Complexity (RCO) reconstruction, Model Driven Prime Editing, PULSE 2.0, Plant optogenetics
Institutions: University of Edinburgh, Heinrich Heine University of Düsseldorf
 https://orcid.org/0000-0002-7975-5013
https://orcid.org/0000-0002-7975-5013
Group Leader in Plant Synthetic Genomics at Heinrich Heine University Düsseldorf and the Cluster of Excellence on Plant Sciences (CEPLAS). My research develops the genetic and cellular methods required for synthetic plant chromosomes to function in living cells — including telomere seeding, neocentromere activation, and heritable gene expression — using Physcomitrium patens as a programmable plant chassis. I use the circadian clock as a quantitative test system to test whether complex gene ...
Projects: CoolWine
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0001-5892-2823
https://orcid.org/0000-0001-5892-2823
Projects: Jahnsen group
Institutions: University of Oslo
Projects: COVID-19 Disease Map
Institutions: National Institute for Infectious Diseases "L.Spallanzani"
Projects: MOSES
Institutions: Medical University Vienna
Projects: COVID-19 Disease Map
Institutions: University of Heidelberg
Projects: Light and plant development
Institutions: University of Edinburgh
Projects: PSYSMO, COVID-19 Disease Map
Institutions: CSIC Madrid
Projects: COVID-19 Disease Map
Institutions: Barcelona Supercomputing Center
Projects: SCaRAB
Institutions: Norwegian University of Science and Technology
Projects: TRANSLUCENT
Institutions: Autonomous University of Barcelona
Projects: SUMO
Institutions: University of Amsterdam
Johan van Beilen University of Amsterdam j.w.a.vanbeilen@uva.nl;
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Wageningen University & Research
 https://orcid.org/0000-0001-9395-5559
https://orcid.org/0000-0001-9395-5559
Projects: MESI-STRAT
Institutions: De Duve Institute
Projects: NAD COMPARTMENTATION
Institutions: University of Bergen
Projects: SulfoSys
Institutions: Wageningen University & Research
I obtained my PhD in 1989 at the Free University (Amsterdam) on a research project in which microbial physiology, biochemistry, and molecular biology were combined. Subsequently I spent 3 years abroad, 2.5 years of which as EMBO fellow at the EMBL (Heidelberg, Germany) where I worked on protein engineering and protein crystallization. I returned to Amsterdam as KNAW fellow for 3 years, during which I worked on protein analysis and pathway engineering. In 1995 I was appointed as group leader ...
Projects: BaCell-SysMO
Institutions: University Medical Centre Groningen
The main area of my expertise concerns protein sorting and secretion in Gram-positive bacteria, such as Bacillus subtilis and Staphylococcus aureus.
The Gram-positive bacterium B. subtilis is well known for its high capacity to secrete proteins into the extracellular milieu, which has led to its exploitation as a "cell factory" for secreted proteins. Nevertheless, the secretion of heterologous proteins of pharmaceutical importance is frequently inefficient. This applied problem has been a major ...
Projects: FAIRDOM user meeting, SYSTERACT, INBioPharm, OXYMOD, BioZEment 2.0
Institutions: University of Leiden, SINTEF
 https://orcid.org/0000-0002-6793-7469
https://orcid.org/0000-0002-6793-7469
POSITION I am an emeritus professor in Biochemistry at the University of Amsterdam (retired 2010).
RESEARCH My research focussed on the human chromatin in its natural environment, i.e. the nucleus of cultured living human cells. Aspects, such as the dynamic folding of the chromatin fiber inside the nucleus and local chemical modification of histones and DNA at genetic loci, are the physical and chemical basis for epigenetic regulation of gene expression. In my group we worked parallel on human ...
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Groningen
I work as a project manager for the Innovative Training Network PoLiMeR - Polymers in the LIver: Metabolism and Regulation funded by the EU. In addition I am a project manager for the UMCG Research BV where I support scientist in the pre-award phase with writing their proposals and in the post-award phase with managing their awarded projects.
Projects: SysMO-LAB
Institutions: University of Amsterdam
Projects: EmPowerPutida
Institutions: Wageningen University & Research
Projects: ROBUSTYEAST
Institutions: VU University Amsterdam
Projects: SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-9143-9726
https://orcid.org/0000-0002-9143-9726
Projects: SysVirDrug
Institutions: Leiden University Medical Center
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: 25TST, 26TOMATO, Sustainable Cacao Genomics Initiative
Institutions: GNV
Projects: NAD COMPARTMENTATION
Institutions: University of Bergen
POSITION Prof. Dr. Natal van Riel is Professor in Computational Modelling at the Academic Medical Center - University of Amsterdam (AMC - UvA) and Associate Professor in Systems Biology and Metabolic Diseases at the Department of Biomedical Engineering of the Eindhoven University of Technology (TU/e).
RESEARCH My research applies mathematical modelling and computation to study metabolic diseases, in particular Metabolic Syndrome and co-morbidities. Systems biology approaches are developed for ...
Projects: COVID-19 Disease Map, SYLOBIO
Institutions: University of Rostock, SBI University of Rostock
 https://orcid.org/0000-0002-2486-0246
https://orcid.org/0000-0002-2486-0246
Projects: SYLOBIO
Institutions: University of Rostock
Projects: NIRBTEST
Institutions: Amsterdam UMC
Projects: Xenophiles Systems Biology
Institutions: VU University Amsterdam
Professor in Planetary Evolution, Department of Earth Sciences, VU Amsterdam Member of Doctorate Committee PhD thesis Joost W Aerts
Projects: COVID-19 Disease Map
Institutions: University of Bonn
Institutions: University of Freiburg
Projects: COVID-19 Disease Map
Institutions: The European Bioinformatics Institute < EMBL-EBI
Projects: Colosys, COVID-19 Disease Map
Institutions: Norwegian University of Science and Technology, Barcelona Supercomputing Center
Projects: ODDITY
Institutions: Universidade de Santiago de Compostela
Projects: Noisy-Strep
Institutions: University of Groningen
The Veening lab is interested in phenotypic bi-stability in Streptococcus pneumoniae and its importance in virulence of this human pathogen.
Projects: COVID-19 Disease Map
Institutions: OROBIX Sarl
Projects: SysMO-LAB
Institutions: University of Heidelberg
Projects: Lipofungi, DeepHyperSpec, Bio4Fuels, LIGNOLIPP
Institutions: Norwegian University of Life Sciences
Projects: SysMetEx
Institutions: Universitity Duisburg-Essen
My research is intended to contribute to the elucidation of the physiological and molecular processes involved in the biofilm formation of acidophilic leaching bacteria with emphasis in their cell-cell communication mechanisms. In SysMetEx, our role is to understand biofilm formation at a microscopical and OMICS levels, in order to optimize it.
Projects: MOSES, SysMO-LAB, BaCell-SysMO
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: SUMO
Institutions: University of Amsterdam
Started out in the field of environmental analytical chemistry and after a few years working in that field, switched to process analysis and chemometrics. Next during my PhD work I came into Life Sciences doing data analysis on microbial batch fermentations (Escherichia coli). During my PhD work my main task was to integrate prior knowledge into data analysis, so called grey modeling. Now my focus lies on white models (based on ordinary differential equations), more specifically building a detailed ...
Projects: FAIRDOM user meeting
Institutions: MDC/BIMSB
Projects: PhotoBoost
Institutions: University of Lancaster
Systems biology for salmon farming is what I do. I lead the DigiSal project (http://tinyurl.com/digisal), whose full title is "Towards the Digital Salmon: From a reactive to a pre-emptive research strategy in aquaculture". DigiSal is part of Digital Life, the first call dedicated to systems biology by the Research Council of Norway. I'm also one of the lead modellers in GenoSysFat (http://tinyurl.com/genosysfat), working to improve the omega-3 content of salmon farmed on sustainable feeds by ...
Projects: DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0009-0003-2141-9652
https://orcid.org/0009-0003-2141-9652
Projects: WP 1: Biodegradable and responsive polymer matrix, WP 2: Antimalarial drug encapsulation, WP 4: Evaluation of drug activity against multiple life cycle stages of malaria parasites, Project Coordination, WP 3: Drug release kinetics study, WP 5: Intradermal 3D drug device administration
Institutions: Leibniz-Institut für Polymerforschung Dresden e. V.
Projects: LEANPROT
Institutions: Competence Center of Food and Fermentation Technologies
Principal Investigator International Clinical Research Center (FNUSA-ICRC) Brno Czech Republic
Projects: COVID-19 Disease Map
Institutions: INSERM
Projects: MS_DILI
Institutions: German Cancer Research Center (DKFZ)
Projects: SCaRAB, BaCell-SysMO
Institutions: University of Greifswald
By training I am a molecular microbiologist with a particular interest in bacterial stress adaptation reactions. Since the early 90ies we have used functional genomics technologies, particularly proteomics, to investigate the response of Bacillus subtilis to environmental challenges. I am currently Head of the Functional Genomics Department of the Interfaculty Institute of Genetics and Functional Genomics at the Ernst-Moritz-Arndt-University Greifswald. Within the Department we are applying ...
Projects: Auromega
Institutions: Norwegian University of Science and Technology
Projects: COSMIC
Institutions: University of Rostock
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: HUMET Startup
Institutions: University of Zurich
Projects: SulfoSys
Institutions: eGene Biotechnologie GmbH
Projects: CropClock
Institutions: Heinrich Heine University of Düsseldorf
Projects: MetApp
Institutions: ETH Zurich
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Goettingen
Projects: SysMO-LAB, NMTrypI - New Medicines for Trypanosomatidic Infections
Institutions: Heidelberg Institute for Theoretical Studies
Group Leader, Molecular and Cellular Modeling Group, EML Research, Heidelberg
Projects: Virtual Human Platform for Safety Assessment
Institutions: Utrecht University
Projects: HUMET Startup
Institutions: Liverpool John Moores University
 https://orcid.org/0000-0003-1916-3822
https://orcid.org/0000-0003-1916-3822
Anton JM Wagenmakers is Professor of Exercise Metabolism and Academic Lead of the Exercise Metabolism and Adaptation Research Group (EMARG) at Liverpool John Moores University, UK. Anton is a leading researcher in skeletal muscle and integrative human physiology and metabolism with 185 peer reviewed journal publications and a Web of Science h-index of 50. His research is focussing on mechanisms by which impairments in fat metabolism lead to impaired glucose tolerance, insulin resistance, and ...
Projects: SBEpo - Systems Biology of Erythropoietin
Institutions: German Cancer Research Center (DKFZ)
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: XyloCut
Institutions: Delft University of Technology (DUT)
 https://orcid.org/0000-0003-2120-1859
https://orcid.org/0000-0003-2120-1859
Assistant professor working on industrial and environmental microorganisms
Projects: ROBUSTYEAST
Institutions: Otto-von-Guericke University Magdeburg
Projects: SulfoSys - Biotec
Institutions: University Duisburg-Essen
Projects: de.NBI-SysBio, GenoSysFat, Kinetics on the move - Workshop 2016, Example use cases, COMBINE Multicellular Modelling, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University of Rostock, University of Greifswald, University Medicine of Greifswald
 https://orcid.org/0000-0002-5886-5563
https://orcid.org/0000-0002-5886-5563
I am a computer scientist by training with a specialisation on database and information systems. Since December 2018 I am professor of Medical Informatics at the University Medicine in Greifswald, Germany, at the Institute of Community Medicine. My lab focuses on research data management in biomedicine, data integration across health care providers, and provenance of clinical research data items within clinical information systems. Furthermore, I am actively involved in COMBINE standardisation ...
Projects: FAIRDOM Community Workers
Institutions: University of Greifswald
Projects: SYSTERACT
Institutions: Chalmers University of Technology
 https://orcid.org/0000-0001-7475-0136
https://orcid.org/0000-0001-7475-0136
Projects: COVID-19 Disease Map
Institutions: University of Pittsburgh
Projects: COVID-19 Disease Map
Institutions: University of California, Los Angeles
Projects: JUANGMC
Institutions: Guizhou Provincial Center for Disease Control and Prevention
 https://orcid.org/0000-0002-7459-2497
https://orcid.org/0000-0002-7459-2497
Projects: TRALAMINOL, CC-TOP
Institutions: University College London (UCL)
Institutions: University of Gothenburg
Projects: HUMET Startup, COVID-19 Disease Map
Institutions: University of Ulster
 https://orcid.org/0000-0002-2750-6410
https://orcid.org/0000-0002-2750-6410
POSITION Lecturer/Assistant Prof in Computational Biology at the University of Ulster.
RESEARCH My research focusses principally on cholesterol metabolism and its role in cardiovascular disease and atherosclerosis. I have also studied its role in innate immunity and infection. In addition, I have worked on areas of personalised and stratified medicine for cardiovascular disease and rheumatoid arthritis.
PUBLICATION My publications can be seen at: https://scholar.google.co.uk/citations?user=oMccxPwAAAAJ&hl=en ...
Projects: Beads analysis
Institutions: Karlsruhe Institute of Technology (KIT)
 https://orcid.org/0000-0002-6151-3999
https://orcid.org/0000-0002-6151-3999
Projects: DigiSal, GenoSysFat, SEEK tutorial for DigiSal
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0002-7450-619X
https://orcid.org/0000-0002-7450-619X
Researcher at the DigiSal project. Recently graduated M.Sc. in Chemistry and Biotechnology.
Projects: Sustainable co-production
Institutions: Leuphana University of Lüneburg
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: University of Gothenburg
Projects: STREAM
Institutions: University of Warwick
Prof. Dr. Volker F. Wendisch is Chair of Genetics of Prokaryotes at the Faculty of Biology at Bielefeld University. Since 2010 he is member of the board of Center for Biotechnology CeBiTec and since 2014 member of the senate of Bielefeld University. He studied biology in Köln University. After having completed his PhD at the Research Center Jülich in 1997, He worked as postdoctoral researcher at the University of California, Berkeley, CA, USA. From 2006-2009 he was Professor for Metabolic Engineering ...
Projects: STREAM, SYSTERACT, INBioPharm, BioZEment 2.0, OXYMOD, BESTER, AquaHealth (ERA-BlueBio)
Institutions: SINTEF
 https://orcid.org/0000-0002-9420-1959
https://orcid.org/0000-0002-9420-1959
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: Sustainable co-production
Institutions: Wageningen University & Research
 https://orcid.org/0000-0002-6009-8543
https://orcid.org/0000-0002-6009-8543
Projects: SysMO-LAB, MOSES, PSYSMO, SulfoSys, SulfoSys - Biotec, EraCoBiotech 2 nd call proposal preparation, Make Me My Model, Mechanism based modeling viral disease ( COVID-19 ) dynamics in human population, Modelling COVID-19 epidemics, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, Xenophiles Systems Biology, Thermodynamics, Non equilibrium thermodynamics, Book on Thermodynamics, and kinetics, Teaching Alien Biology, Outdated material, Fusion-fission-mitophagy, Stochastics and bursting, Thermotables for biochemistry, Thermotables
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester, VU University Amsterdam, University of Amsterdam, Systems Biology Amsterdam
 https://orcid.org/0000-0002-0443-6114
https://orcid.org/0000-0002-0443-6114
Systems Biologist at University of Amsterdam, Free University Amsterdam, University of Manchester, Infrastructure Systems Biology.NL (ISBE.NL), Systems Biology Amsterdam.
Projects: MultiDefence - sLoLa
Institutions: University of Exeter
 https://orcid.org/0000-0003-4396-0354
https://orcid.org/0000-0003-4396-0354
Projects: WURSynBio
Institutions: Wageningen University & Research
Projects: SYSTERACT
Institutions: University of Leiden
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: SulfoSys
Institutions: University of Braunschweig
Projects: MIX-UP
Institutions: Forschungszentrum Jülich
Projects: COVID-19 Disease Map
Institutions: Imperial College London
Projects: STREAM
Institutions: University of Warwick
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
 https://orcid.org/0000-0003-3954-8981
https://orcid.org/0000-0003-3954-8981
Projects: MOSES
Institutions: Manchester Centre for Integrative Systems Biology, University of Manchester
Projects: HUMET Startup
Institutions: Leiden University Medical Center
 https://orcid.org/0000-0002-2172-7394
https://orcid.org/0000-0002-2172-7394
Professor in genetics and systems biology of the metabolic syndrome
Projects: COVID-19 Disease Map
Institutions: US Environmental Protection Agency
Projects: COVID-19 Disease Map, VHP project
Institutions: University Maastricht
Projects: SysMetEx
Institutions: Systems Biomedicine Center, University of Luxembourg
Projects: Coastal Data
Institutions: Afbi
Projects: SEEK tutorial for DigiSal, DigiSal
Institutions: Norwegian University of Science and Technology, Norwegian University of Life Sciences
Projects: COSMIC
Institutions: University of Nottingham
A microbiologist with interest in microbial physiology and metabolism. Current research focuses on the biology of pathogenic and non-pathogenic clostridia, in particluar in vivo metabolism and cell-to-cell communication. Also active in metabolic engineering, using both systems and synthetic biology approaches.
Projects: COVID-19 Disease Map
Institutions: University of Newcastle
Projects: de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, FAIRDOM, EnzymeML, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", FAIRDOM Community Workers, MIX-UP, COVID-19 Disease Map, ERNEST Mapping Group Pilot Study, NMTrypI - New Medicines for Trypanosomatidic Infections, Standardization of enzyme-catalyzed reaction measurement, Standardization of enzyme-catalyzed reaction modelling, ModeleXchange initiative, CoVIDD - Coronavirus interactions in drug discovery - optimization and implementation, Mass spectrometry proteomics for biomarker discovery, Thymidylate synthase dimer dissociation, WG1 - Compound libraries coordination and integration of compound design, WG2 - Integration of early phase studies and low environmental impact actions, WG3 - Coordination of in vitro-to-in vivo translation of OneHealth leads and candidates, WG4 - Integration of R&D process-environmental studies and translation in informed whitepaper, WG5 - Promote dissemination, WG6 - Promote the transfer of knowledge, SABIO-VIS, SDBV/HITS, BioTarget Database, Compound Data
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-9077-5664
https://orcid.org/0000-0002-9077-5664
Projects: BaCell-SysMO
Institutions: University of Braunschweig
University Education: 1987-1993, Biotechnology (Diploma), Technische Universität Braunschweig, Germany.
Dissertation: 1993-1996, Disseration German Research Centre for Biotechnology, Biochemical Engineering Division, Braunschweig, Germany.
Habiliation: 2006, Saarland University, Saarbrücken, Germany.
Positions 1993-1996: Research Assistant, German Research Centre for Biotechnology, Biochemical Engineering Division, Braunschweig, Germany. 1997-1998: Post-doc at Department of Applied Chemistry & ...
Projects: HUMET Startup, COMBINE Multicellular Modelling
Institutions: Humboldt University Berlin, Humboldt-Universität zu Berlin
Projects: EbN1 Systems Biology, Heat stress response of the red-tide dinoflagellate Prorocentrum cordatum, The nucleus of Prorocentrum cordatum, Conspicuous chloroplast with LHC‒PSI/II‒megacomplex and diverse PBPs in the marine dinoflagellate Prorocentrum cordatum, Proteogneomics of Desulfobacteraceae members
Institutions: Carl von Ossietzky University of Oldenburg Institute for Chemistry and Biology of the Marine Environment (ICBM)
 https://orcid.org/0000-0001-7253-3373
https://orcid.org/0000-0001-7253-3373
Projects: Group Data Science, Group Microbial Interface Biology, Core Facility Fluorescence Cytometry, Targeting the immunoproteasome: Analyzing the biological effects of specific versus pan inhibition, Macrophage and neutrophil redox proteome in infection with Mycobacterium tuberculosis, Methylation differences during respiratory infection, Lipid droplet formation and dynamics in murine macrophages, Implementation of Nanopore Sequencing for Detection of Treatment Induced Transcriptomic and Epitranscriptomic Changes in Leukaemic Tumour Models, Revisiting mutational resistance to ampicillin and cefotaxime in Haemophilus influenzae, OMICS of vascular smooth muscle cells under oxidative stress, Systems genetics of lung health and asthma (GWAS, PGS, QTL), Archaean stimulation of human primary PBMCs and PBMCs monocytes, iMED-RISK: Improvement of risk stratification in infant medulloblastoma using integrated models of clinical, histopathological and molecular data
Institutions: Forschungszentrum Borstel
 https://orcid.org/0000-0003-4004-0464
https://orcid.org/0000-0003-4004-0464
Projects: SulfoSys - Biotec
Institutions: Sigma Aldrich ( Swiss)
Institutions: University of Tuebingen, University of Tübingen
Projects: MOSES, ExtremoPharm
Institutions: University of Heidelberg
Projects: COSMIC, BaCell-SysMO, SYSTERACT, HUMET Startup, INCOME, BESTER, iRhythmics, COVID-19 Disease Map, SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke, Implementation of Nanopore Sequencing for Detection of Treatment Induced Transcriptomic and Epitranscriptomic Changes in Leukaemic Tumour Models
Institutions: University of Rostock
I am an Assistant Professor at Leiden University in the Leiden Institute of Advanced Computer Science. I am a bioinformatician and my research interests are in data integration. I use scientific workflows and semantic web technologies to integrate and analyse data in systems biology and functional genomics.
Projects: MIX-UP
Institutions: RWTH Aachen University
Projects: TestingSeek, FAIRDOM Community Workers, FAIRDOM
Institutions: University of Edinburgh
Projects: Selective Destruction in Ageing
Institutions: University of Newcastle
Bioinformatician / Computational Biologist
Projects: SulfoSys, SulfoSys - Biotec
Institutions: University of Sheffield
I am the foundation Professor of Systems Biology and Engineering within the Department of Chemical and Process Engineering (CPE), at The University of Sheffield. My research philosophy is centred on a mechanistic systems biology approach to solve biochemical reaction engineered processes. I wish to pursue issues involved in the effective utilisation of biological resources. The approach is specifically targeted at the conjunction of chemical engineering (metabolic engineering and synthetic biology), ...
Projects: BaCell-SysMO
Institutions: DSM Nutritional Products Ltd.
Projects: HubMOL
Institutions: University Medical Center Groningen
 https://orcid.org/0009-0004-5666-1032
https://orcid.org/0009-0004-5666-1032
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: KTH Royal Institute of Technology
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Radboud University Nijmegen
I am a first year graduate student in the lab of Prof. Wilhelm Huck at the Radboud University Nijmegen. I am working on a project to create an artificial cell. This involves implementing complex genetic networks in cell-free systems which are far from equilibrium. My project involves designing, quantifying, modeling such networks. I am a molecular biologist by training and have basic modeling skills. I am looking to expand my skill-set to include microscopy, microfluidics and a little bit of ...
Projects: DCM
Institutions: Institute of Cytology and Genetics
 https://orcid.org/0000-0002-0712-6939
https://orcid.org/0000-0002-0712-6939
Projects: MIX-UP
Institutions: Tsinghua University
Projects: Matsutake
Institutions: Seoul National Univ.
Projects: LabscriptAI
Institutions: University of Chinese Academy of Sciences
Projects: COVID-19 Disease Map
Institutions: Elsevier
Projects: Noisy-Strep
Institutions: University of Newcastle
Projects: pISA-tree, HYp - Spatiotemporal analysis of hypersensitive response to Potato virus Y in potato, INDIE - Biotechnological production of sustainable indole, _p_stRT, ADAPT - Accelerated Development of multiple-stress tolerAnt PoTato, tst, tst2
Institutions: National Institute of Biology, tst
 https://orcid.org/0000-0002-1669-6482
https://orcid.org/0000-0002-1669-6482
Computational Biologist and Biostatistician at Department of Biotechnology and Systems Biology, National Institute of Biology (NIB)
Projects: FAIRe Tests BMDS
Institutions: Faculty of Medicine, Martin Luther University Halle-Wittenberg
Projects: TRANSLUCENT
Institutions: Institute of Physiology Academy of Sciences
Projects: MOSES, ExtremoPharm, ZucAt, GenoSysFat, DigiSal, EraCoBiotech 2 nd call proposal preparation, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: University of Stuttgart, University of Hohenheim, Norwegian University of Life Sciences, Norwegian University of Science and Technology
 https://orcid.org/0000-0002-7973-9902
https://orcid.org/0000-0002-7973-9902
I've become a SysMO DB PAL for MOSES project in 2007 being a post-doc in lab of Prof. Matthias Reuss at University of Stuttgart. In the MOSES project, our major efforts were in the experimental data acquisition for dynamic model of primary carbon and anaerobic energy metabolism in yeast. The model implements prediction of perturbations of two types: glucose pulse and temperature jump. We implement “stimulus-response” methodology for the unraveling the dynamic structure of the network and to ...
Projects: FAIRDOM user meeting, Systems toxicology of Atlantic cod, DigiSal, GenoSysFat, FAIRDOM Community Workers
Institutions: University of Bergen
Projects: SulfoSys
Institutions: University Duisburg-Essen
Projects: BaCell-SysMO
Institutions: University of Marburg
Projects: Millar group
Institutions: University of Edinburgh
Projects: BioZEment 2.0
Institutions: Norwegian University of Science and Technology
Projects: Noisy-Strep
Institutions: University of Newcastle
Projects: SysMO-LAB
Institutions: University of Rostock
Projects: XyloCut
Institutions: Delft University of Technology (DUT)
Projects: Systems toxicology of Atlantic cod
Institutions: University of Bergen
 https://orcid.org/0000-0003-4684-317X
https://orcid.org/0000-0003-4684-317X
Projects: COSMIC
Institutions: University of Nottingham
Projects: BIOMICRO PROJECT
Institutions: MIT
Projects: FAIRDOM user meeting, PlaSMo model repository
Institutions: University of Edinburgh
Projects: KOSMOBAC
Institutions: Ludwig-Maximilians University of Munich
Projects: Lipofungi, Bio4Fuels
Institutions: Norwegian University of Life Sciences
 https://orcid.org/0000-0001-5046-520X
https://orcid.org/0000-0001-5046-520X
Projects: SysVirDrug
Institutions: University of Greifswald
Projects: NTNU Health Druglogics, Colosys
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-3609-8674
https://orcid.org/0000-0002-3609-8674
Projects: HUMET Startup
Institutions: Wageningen University & Research
I am a researcher (PhD student) working at Wageningen University & Research as bioinformatician and modeller. I am working as part of the MycoSynVac (http://www.mycosynvac.eu/) project on dynamic modelling of central carbon metabolism in M. pneumoniae, to be extended to full dynamic modelling of metabolism to be implemented in a whole cell model. I am also looking into possibilities to improve standards in model generation using semantic technologies, improving automatic generation, annotation ...
Projects: COVID-19 Disease Map
Institutions: Pacific Northwest National Laboratory
Projects: test
Institutions: Centre national de la Recherche Scientifique (CNRS)
Projects: LiceVault
Institutions: University of Bergen
Projects: Hi-IMPAcTB
Institutions: University of Toronto