Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0003-3634-190X
https://orcid.org/0000-0003-3634-190X
Expertise: Computational Biology, Mathematical modelling, Multicellular Modelling, Compucell3D, Virtual Tissues, Model Sharing, Language Standards, Agent-based modelling, Dynamic modelling, Computational Systems Biology, standards, Developmental Biology, Toxicology, Framework Development, Cancer, Immunology, Community Building
Tools: Compucell3D, Antimony, tellurium, SBML, CC3DML
Dr. Glazier’s research focuses on early embryonic development, developmental and chronic toxicity and disease, with more than 100 experimental and computational papers on biological development and developmental diseases (including polycystic kidney disease (ADPKD), tumor growth and vascularization, Age Related Macular Degeneration and diabetic retinopathies, somitogenesis and liver toxicity) and more recently on modeling in-host viral infection and immune response. As part of his work on infection ...
Projects: COMBINE Multicellular Modelling
Institutions: University of Florida
 https://orcid.org/0000-0002-4274-656X
https://orcid.org/0000-0002-4274-656X
Expertise: Computational Biology, Biophysics, Biological Simulation
Tools: Compucell3D, Tissue Forge, libRoadrunner, SBML, Antimony, MaBoSS
Projects: Simulation Foundries, CML for thermophysical properties of mixtures, Test Project (dummy), Towards Reproducible Enzyme Modeling
Institutions: University of Stuttgart, Karlsruhe Institute of Technology (KIT)
 https://orcid.org/0000-0001-9119-1778
https://orcid.org/0000-0001-9119-1778
Expertise: enzyme kinetics, enzymes, Enzymatic reactions, biochemical enzyme characterization, Biochemistry, molecular simulation, molecular modeling, Programming, Bioinformatics, Computational Biology
Tools: Gromacs, Python, Molecular Dynamics, bash, Biochemistry, Bioinformatics, Biochemistry and protein analysis, Enzyme assay, enzyme kinetics, isothermal titration calorimetry, dynamic light scattering, Spectrophotometry
Polyglot European Scientist. I thrive working in interdisciplinary environments combining the study of enzyme reactions and mechanisms with bioinformatics, molecular modelling, automated data analysis and data stewardship.
Projects: pISA-tree, HYp - Spatiotemporal analysis of hypersensitive response to Potato virus Y in potato, INDIE - Biotechnological production of sustainable indole, _p_stRT, ADAPT - Accelerated Development of multiple-stress tolerAnt PoTato, tst, tst2
Institutions: National Institute of Biology, tst
 https://orcid.org/0000-0002-1669-6482
https://orcid.org/0000-0002-1669-6482
Expertise: Molecular Biology, Statistics, Bioinformatics, Mathematical and statistical modeling, Programming, Data analysisMathematical modellingBioinformaticsSystems biology, Data Management, Data analysis, Visualization, Data Integration, Computational Biology
Tools: Bioinformatics, Computational and theoretical biology, Computational Systems Biology, Data Management, Databases, Dynamic modelling, Molecular Biology, Python, R, Systems Biology, Data Integration
Computational Biologist and Biostatistician at Department of Biotechnology and Systems Biology, National Institute of Biology (NIB)
Projects: NTNU Health Druglogics, Colosys
Institutions: Norwegian University of Science and Technology
 https://orcid.org/0000-0002-3609-8674
https://orcid.org/0000-0002-3609-8674
Expertise: Systems Biology, Computational Biology
Projects: HUMET Startup, COVID-19 Disease Map
Institutions: University of Ulster
 https://orcid.org/0000-0002-2750-6410
https://orcid.org/0000-0002-2750-6410
POSITION Lecturer/Assistant Prof in Computational Biology at the University of Ulster.
RESEARCH My research focusses principally on cholesterol metabolism and its role in cardiovascular disease and atherosclerosis. I have also studied its role in innate immunity and infection. In addition, I have worked on areas of personalised and stratified medicine for cardiovascular disease and rheumatoid arthritis.
PUBLICATION My publications can be seen at: https://scholar.google.co.uk/citations?user=oMccxPwAAAAJ&hl=en ...
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
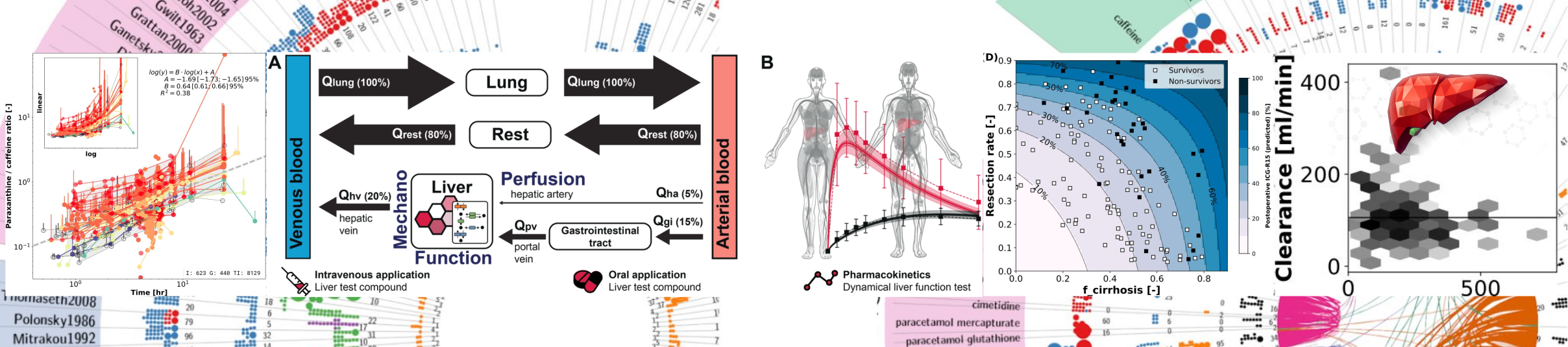
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: ZucAt
Institutions: Institute of Cytology and Genetics, Novosibirsk State University
Expertise: computational genomics, Computational Biology
Tools: high-throughput sequencing, ChIP-seq
I am Senior Scientist at Institute of Cytology and Genetics SB RAS in Novosibirsk. My research focus is computational genomics: high-throughput sequencing, ChIP-seq, human genome research (HUGO member), high performance computing in bioinformatics
Projects: SUMO
Institutions: University of Sheffield
Professor of Computer Science, University of Sheffield. FBCS, FIMA, CEng, C.Math, CITP. I have been involved in the use of computational techniques for modelling biological systems since 1980. More recently I have developed a technique of agent-based modelling based on the framework FLAME which is the only such system that can be run on supercomputers. We have made significant new biological discoveries using this approach: The approach models the location and activity of millions of individual ...