Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0003-3634-190X
https://orcid.org/0000-0003-3634-190X
Expertise: Computational Biology, Mathematical modelling, Multicellular Modelling, Compucell3D, Virtual Tissues, Model Sharing, Language Standards, Agent-based modelling, Dynamic modelling, Computational Systems Biology, standards, Developmental Biology, Toxicology, Framework Development, Cancer, Immunology, Community Building
Tools: Compucell3D, Antimony, tellurium, SBML, CC3DML
Dr. Glazier’s research focuses on early embryonic development, developmental and chronic toxicity and disease, with more than 100 experimental and computational papers on biological development and developmental diseases (including polycystic kidney disease (ADPKD), tumor growth and vascularization, Age Related Macular Degeneration and diabetic retinopathies, somitogenesis and liver toxicity) and more recently on modeling in-host viral infection and immune response. As part of his work on infection ...
Projects: COMBINE Multicellular Modelling
Institutions: University of Florida
 https://orcid.org/0000-0002-4274-656X
https://orcid.org/0000-0002-4274-656X
Expertise: Computational Biology, Biophysics, Biological Simulation
Tools: Compucell3D, Tissue Forge, libRoadrunner, SBML, Antimony, MaBoSS
Rahuman Sheriff is a Senior Project Leader (BioModels) at the European Bioinformatics Institute, European Molecular Biology Laboratory (EMBL-EBI), Hinxton, Cambridge, UK. He interests include mathematical modelling, development of novel tools and resources for building models, immune digital twin, quantitative imaging, single cell systems biology and chemoinformatics. He is one of the editors of Systems Biology Markup Language (SBML).
Projects: COVID-19 Disease Map
Institutions: National Institute of Informatics
 https://orcid.org/0000-0001-8725-3366
https://orcid.org/0000-0001-8725-3366
Expertise: Bioinformatics, Computational Systems Biology, Curation, Data analysis, Systems Biology
Tools: SBML, Python, Matlab, Systems Biology, Bioinformatics
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0002-0248-6679
https://orcid.org/0000-0002-0248-6679
Projects: COVID-19 Disease Map
Institutions: independent researcher
 https://orcid.org/0000-0002-3882-6073
https://orcid.org/0000-0002-3882-6073
Expertise: Data analysis, Bioinformatics, Molecular Biology, Genetics, Neuroscience
Tools: Data Science, R, Cell designer, SBML, Single Cell analysis, Transcriptomics
Projects: COVID-19 Disease Map, Covid-19 Interferon pathway modelling and analysis, Boolean modeling of Parkinson disease map
Institutions: Luxembourg Centre for Systems Biomedicine (LCSB), University of Luxembourg
 https://orcid.org/0000-0001-7403-181X
https://orcid.org/0000-0001-7403-181X
Expertise: Machine Learning, Deep Learning, Modeling, Medical microbiology, Molecular Biology
Tools: Python, R, Matlab, Javascript, SBML, Dynamic modelling, microbiology techniques, Shell scripting
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0003-3851-9978
https://orcid.org/0000-0003-3851-9978
Expertise: Curation, Computational Systems Biology, Constraint-based Modelling, Systems Biology
Tools: SBML, Python, cobrapy toolbox, CellDesigner, libSBML, jupyter notebooks
Projects: COVID-19 Disease Map
Institutions: University of Tübingen
 https://orcid.org/0000-0002-1240-5553
https://orcid.org/0000-0002-1240-5553
Expertise: Systems Biology, Computational Systems Biology, Databases, Dynamic modelling, Java, Mathematical modelling, Metabolic Engineering, Disease Maps, Curation, Modeling, Data Integration, Constraint-based Modelling, Parameter estimation
Tools: SBML, SBGN, SBGNML, JSBML, Jupyter, Python, cobrapy toolbox, SBSCL, InSilico, Kinetic Modeling
Andreas Dräger is the assistant professor for Computational Systems Biology of Infection and Antimicrobial-Resistant Pathogens at the University of Tübingen in Germany. His group aims to combat the spreading antibiotics resistances by using mathematical modeling and computer simulation of bacterial systems up to entire microbiomes and host-pathogen interactions. In doing so, his group actively contributes to the advancement of various COMBINE standards.
Expertise: Systems Biology, SBML, Java, Python, Machine Learning, Mathematical modelling, SBGN, Curation
Tools: CellDesigner, SBML, SBGN
Projects: Kinetics on the move - Workshop 2016, COVID-19 Disease Map
Institutions: Kinetics on the move Workshop at HITS, Luxembourg Centre for Systems Biomedicine (LCSB)
 https://orcid.org/0000-0003-1473-370X
https://orcid.org/0000-0003-1473-370X
Expertise: Data Management, Molecular Biology, Systems Biology, Curation
Tools: SBGN, CellDesigner, MIRIAM, SBML, Data Science
Scientific Project Manager at Luxembourg Centre for Systems Biomedicine, University of Luxembourg http://lcsb.uni.lu
Projects: COVID-19 Disease Map
Institutions: Barcelona Supercomputing Center
 https://orcid.org/0000-0002-7696-1241
https://orcid.org/0000-0002-7696-1241
I completed my BSc in Biology and MSc in Cell Biology by the University of Valencia. During my last undergrad year I participated in synthetic biology’s iGEM competition where I dove in the use of models in Biology, which pushed me to pursue a PhD in the Department of Applied Mathematics in the Technical University of Valencia.
My research on Metabolic Engineering of hydrogen in cyanobacteria led me to be visiting researcher at Uppsala University, Denmark Technical University and EMBL Heidelberg. ...
Expertise: Biochemistry, Cell biology, Data analysis, Dynamic modelling, Systems Biology, Image analysis, Genetics, Molecular Biology, R, SBML, Curation, Quantitative Biology, Physical Chemistry
Tools: Biochemistry and protein analysis, Bioinformatics, Systems Biology, SBML, R, ODE, Molecular biology techniques (RNA/DNA/Protein), Genetics, Dynamic modelling, Computational and theoretical biology, CellDesigner, Parameter estimation
Projects: COVID-19 Disease Map
Institutions: Integrative BioInformatics Inc.
 https://orcid.org/0000-0001-7979-2386
https://orcid.org/0000-0001-7979-2386
Expertise: Cell physiology, Cell biology, ODE modeling, Human physiology, Kinetics, Tracer Kinetics
Location: Mountain View, CA, USA
Institutions: Latvia University of Agriculture
Expertise: Python, Systems Biology, Dynamic modelling, Mathematical modelling
Tools: COBRA toolbox, cobrapy toolbox, Python, SBML, Copasi, Computational Systems Biology
Projects: Millar group, TiMet, PHYTOCAL: Phytochrome Control of Resource Allocation and Growth in Arabidopsis and in Brassicaceae crops, POP - the Parameter Optimisation Problem, Regulation of flowering time in natural conditions, PlaSMo model repository
Institutions: University of Edinburgh
 https://orcid.org/0000-0003-1756-3654
https://orcid.org/0000-0003-1756-3654
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0002-5901-1404
https://orcid.org/0000-0002-5901-1404
I am a multicell and multiscale modeler and member of the Biocomplexity group at Indiana University. The group produces and maintains the multicellular modelling package Compucell3D. I am also a voting member of the USA committee to the International Organization for Standards (ISO) working group on standards in biotechnology and the chairman of a sub-sub group on publishing standards.
Projects: COMBINE Multicellular Modelling
Institutions: University College London (UCL)
 https://orcid.org/0000-0001-5963-8576
https://orcid.org/0000-0001-5963-8576
Expertise: Dynamic modelling, Databases, Mathematical modelling, standards, Neuroscience, NeuroML
Biochemist currently keeping busy as: Research Data Manager (Vrije Universiteit Amsterdam), Software Engineer (Heidelberg University) and member of the SBML Development Team (Caltech).
Lutz Brusch is heading the research group "Spatio-temporal pattern formation in cells and tissues" at the Centre for Information Services and High Performance Computing of TU Dresden, Germany. The group is co-developing the multi-cellular modelling and simulation framework Morpheus (https://morpheus.gitlab.io) and is collaborating with experimental labs on questions of tissue morphogenesis and regeneration.
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
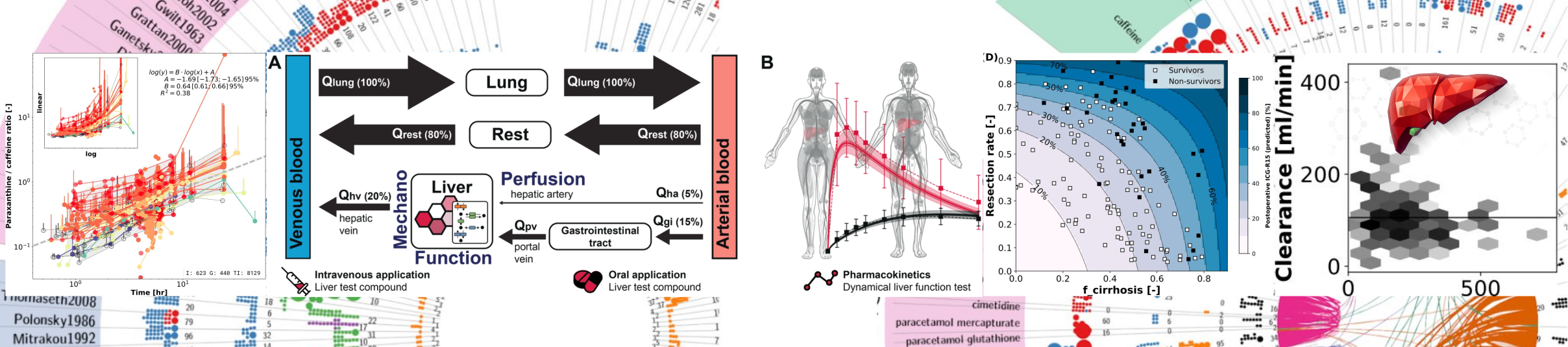
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: de.NBI-SysBio, GenoSysFat, Kinetics on the move - Workshop 2016, Example use cases, COMBINE Multicellular Modelling, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University of Rostock, University of Greifswald, University Medicine of Greifswald
 https://orcid.org/0000-0002-5886-5563
https://orcid.org/0000-0002-5886-5563
I am a computer scientist by training with a specialisation on database and information systems. Since December 2018 I am professor of Medical Informatics at the University Medicine in Greifswald, Germany, at the Institute of Community Medicine. My lab focuses on research data management in biomedicine, data integration across health care providers, and provenance of clinical research data items within clinical information systems. Furthermore, I am actively involved in COMBINE standardisation ...
Projects: SysMO-LAB
Institutions: University of Heidelberg
I am working on a kinetic model of the central metabolism as well as on a genome wide model of Streptococcus pyogenes.
Projects: SUMO
Institutions: University of Stuttgart
Expertise: Mathematical modelling, Data Management, Systems Biology, Parameter estimation
Tools: SBML, ODE, Matlab, Mathematica
Former: PhD student as research associate at the Institute for System Dynamics (ISYS), Universität Stuttgart, Germany. Engineering background→modelling, identification and analyses. Detailed kinetic modelling, identification and analysis of the TCA cycle (tricarboxylic acid cycle, citric acid cycle) and the ETC (electron transport chains, respiratory chains) of Escherichia coli. One of the SysMO-DB pals for SUMO. Now: Industrial affiliation
Projects: COSMIC
Institutions: Beuth University of Applied Sciences Berlin
 https://orcid.org/0000-0001-6096-1354
https://orcid.org/0000-0001-6096-1354
Expertise: Mathematical modelling, Data Management, coupling metabolome and environome, rapid sampling experiments, dynamics of biological networks, bioreactor models, Optimal experimental design, Dynamic optimization., Nonlinear Dynamics, Data Integration, Parameter estimation
Tools: SBML, Matlab, Fermentation, Material balance based modeling, stimulus response experiments, evaluation of process dynamics, continuous cultivation, Dynamic modelling
Process engineer, modeling biological systems since 1985.
Projects: STREAM
Institutions: University of Warwick
Systems Biologist specialising in data integration, high-throughput sequence analysis, and evolutionary and comparative analyses.
Projects: COSMIC
Institutions: University of Rostock
biomathematician, PhD student at the University of Rostock, Systems Biology Group Rostock
Projects: TRANSLUCENT
Institutions: Humboldt-Universität zu Berlin
Expertise: Bioinformatics, Data Management
I created this for all SysMo Modellers
http://www.semanticsbml.org/aym Annotate Your Model
There you can annotate your non SBML models with biological terms (MIRIAM annotations). As a cool extra you can view you model source code with inserted biological infomation.
Together with this http://www.semanticsbml.org/semanticSBML you can serach for similar BioModels. The similarity search is based on MIRIAM annotations that are attached to you model. AYM also allows you to create annotations without ...
Expertise: Molecular Biology, Bioinformatics, Mathematical modelling, Reactor models, dynamics of biological networks, Mathematical and statistical modeling, bioreactor models, Dynamics and Control of Biological Networks, Parameter estimation
Tools: Bioinformatics, Computational and theoretical biology, Transcriptomics, Model organisms, Single Cell analysis, SBML, ODE, Linear equations, Matlab, Microarray analysis, linux, Material balance based modeling, stimulus response experiments, DIVA, differential algebraic equations, evaluation of process dynamics, continuous cultivation
I'm an engineer at the MPI Magdeburg and I'm working in the field of mathematical modeling, model verification, parameter identification, model analysis and experimental design. I'm involved in two projects, KOsmoBac and PSYSMO.
Projects: KOSMOBAC
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: SysMO-LAB
Institutions: Wageningen University & Research
Expertise: Bioinformatics, Mathematical modelling, Reactor models, dynamics of biological networks.
Tools: Molecular Biology, Cell biology, Computational and theoretical biology, Metabolomics, Model organisms, SBML, ODE, Partial differential equations, Algebraic equations, Linear equations, Copasi, JWS Online, Matlab, Mathematica, SQL, Material balance based modeling
I'm a modeller, specialized in kinetic modeling of biochemical networks. My focus in the SysMO-LAB consortium is on creating models of Lactococcus lactis glycolysis and couple this to other related lactic acid bacteria like Streptococcus pyogenes and Enterococcus faecalis. Besides kinetic modeling, I'm also interested in combining various modeling techniques (genome-scale modeling, qualitative modeling).