Projects: MESI-STRAT, NAD COMPARTMENTATION, NAD phylogeny, HubMOL
Institutions: The Arctic University of Norway (UiT)
 https://orcid.org/0000-0002-9124-5112
https://orcid.org/0000-0002-9124-5112
Rahuman Sheriff is a Senior Project Leader (BioModels) at the European Bioinformatics Institute, European Molecular Biology Laboratory (EMBL-EBI), Hinxton, Cambridge, UK. He interests include mathematical modelling, development of novel tools and resources for building models, immune digital twin, quantitative imaging, single cell systems biology and chemoinformatics. He is one of the editors of Systems Biology Markup Language (SBML).
Projects: COVID-19 Disease Map
Institutions: University of Surrey
 https://orcid.org/0000-0001-5640-7422
https://orcid.org/0000-0001-5640-7422
Reader (Professor) of Systems Biology; Executive Director for the International Society of Systems Biology (ISSB); Editor-in-Chief of Current Opinion in Systems Biology (Elsevier).
Expertise: Biochemistry, Cell biology, Data analysis, Dynamic modelling, Systems Biology, Image analysis, Genetics, Molecular Biology, R, SBML, Curation, Quantitative Biology, Physical Chemistry
Tools: Biochemistry and protein analysis, Bioinformatics, Systems Biology, SBML, R, ODE, Molecular biology techniques (RNA/DNA/Protein), Genetics, Dynamic modelling, Computational and theoretical biology, CellDesigner, Parameter estimation
Projects: PoLiMeR - Polymers in the Liver: Metabolism and Regulation
Institutions: University of Groningen
I work as a project manager for the Innovative Training Network PoLiMeR - Polymers in the LIver: Metabolism and Regulation funded by the EU. In addition I am a project manager for the UMCG Research BV where I support scientist in the pre-award phase with writing their proposals and in the post-award phase with managing their awarded projects.
Expertise: Mathematical modelling, Systems Biology, Parameter estimation
Tools: Dynamic modelling, Matlab, ODE, Parameter estimation
Projects: INCOME, COVID-19 Disease Map
Institutions: ICB Helmholtz Center Munich, University of Bonn
 https://orcid.org/0000-0002-4935-3312
https://orcid.org/0000-0002-4935-3312
Expertise: Mathematical modelling, Systems Biology, Parameter estimation
I studied Engineering Cybernetics at the University of Stuttgart and the University of Wisconsin, Madison. After my graduation, I started my PhD studies in systems biology for which I received a Ph.D. degree in 2013. A few months later I became team leader at the Institute of Computational Biology at the Helmholtz Zentrum München. Since August 2015, I lead an independent junior research group at the Helmholtz Zentrum München.
My research focuses on the development of methods for the data-driven ...
I am had of the Research Group PiDOMICS which aims at the identification of human biomarkers for fungal infection using omics-data. Moreover, I am PI Infrastructure project of the Collaborative Research Center / Transregio 124 Pathogenic fungi and their human host: Networks of Interaction - FungiNet. Thereby my expertise is the implementation and usage of pipeline for OMICS (genome, transcriptome, protoem) data analysis, as well as data-warehouses for visualizing these data.
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
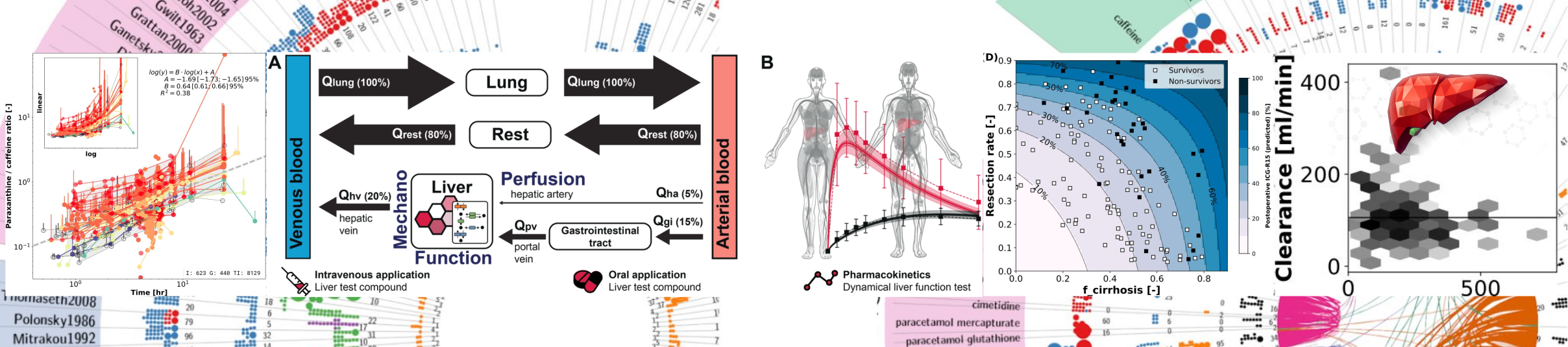
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: MOSES, ExtremoPharm, ZucAt, GenoSysFat, DigiSal, EraCoBiotech 2 nd call proposal preparation, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: University of Stuttgart, University of Hohenheim, Norwegian University of Life Sciences, Norwegian University of Science and Technology
 https://orcid.org/0000-0002-7973-9902
https://orcid.org/0000-0002-7973-9902
Expertise: Biochemistry, coupling metabolome and environome, rapid sampling experiments, Systems Biology, carbon metabolism, Stoichiometric modelling, Proteomics, Metabolomics, yeast, fungi, Dynamics and Control of Biological Networks
Tools: Biochemistry and protein analysis, Metabolomics, Matlab, Fermentation, Chromatography, Material balance based modeling, stimulus response experiments, continuous cultivation, Enzyme assay, Mass spectrometry (LC-MS/MS), HPLC, GC and LC/MS analysis of metabolites, ODE, Parameter estimation
I've become a SysMO DB PAL for MOSES project in 2007 being a post-doc in lab of Prof. Matthias Reuss at University of Stuttgart. In the MOSES project, our major efforts were in the experimental data acquisition for dynamic model of primary carbon and anaerobic energy metabolism in yeast. The model implements prediction of perturbations of two types: glucose pulse and temperature jump. We implement “stimulus-response” methodology for the unraveling the dynamic structure of the network and to ...
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Åbo Akademi University
Projects: SulfoSys - Biotec, ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Otto-von-Guericke University Magdeburg, University of Gothenburg
 https://orcid.org/0000-0001-6971-2530
https://orcid.org/0000-0001-6971-2530
Expertise: Signalling networks, dynamics of biological networks., Databases, Data analysis, Systems Biology, Model selection, Identifiability, Cellular Senescence, Cell Cycle, Dynamic Systems, Image processing, Image analysis, Parameter estimation
Tools: ODE, FACS, Model selection, Fluorescence and confocal microscopy, Identifiability analysis, Parameter estimation
My group investigates dynamic regulation and control mechanisms of cellular signal transduction networks by a combination of theoretical, experimental and computational methods. We seek to make sense of our biological data with the help of mathematical models, which ideally enable us to make valid predictions for new experiments, thereby generating novel biological insights.
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Institute of Cytology and Genetics
I am a biomodeler, PhD student. Actually, I've graduated from Novosibirsk State University on two specialities: my bachelor diploma is done in computer science and the master thesis is defended in information biology. So, I'm kind of drifting towards biology. I am a part of the Haploid Evolutionary Constructor project. Our research group studies are dedicated to the simulation of prokaryotic communities. Personally, I am involved into the simulation of spatially distributed bacterial communities ...
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Universität Konstanz
Expertise: Molecular Biology, Systems Biology, Mathematical modelling
Tools: quantitative western blot analysis, Molecular Biology, Java, octave, R, ODE
I studied Life Science (which is similar to chemical biology) at the University of Konstanz and became interested in Bioinformatics, Systems Biology and quantitative analyses during my Master's. In my PhD project I combine experimental analyses with modeling and parameter estimation approaches to quantitatively analyse regulation of apoptosis at the level of the Bcl-2 protein family.
Projects: ICYSB 2015 - International Practical Course in Systems Biology
Institutions: Imperial College London
I am a PhD student in the Theoretical Systems Biology group, based at Imperial College London. The aim of my PhD is to understand how noise can be the driving force of decision-making processes (differentiation, self-renewal, apoptosis or tumorgenesis), and what are our chances to control them. So far I have been working on method development for stochastic models, a moment closure framework and a stochastic reachability method, to look into cell-to-cell-variability.
Expertise: Molecular Biology, Bioinformatics, Mathematical modelling, Reactor models, dynamics of biological networks, Mathematical and statistical modeling, bioreactor models, Dynamics and Control of Biological Networks, Parameter estimation
Tools: Bioinformatics, Computational and theoretical biology, Transcriptomics, Model organisms, Single Cell analysis, SBML, ODE, Linear equations, Matlab, Microarray analysis, linux, Material balance based modeling, stimulus response experiments, DIVA, differential algebraic equations, evaluation of process dynamics, continuous cultivation
I'm an engineer at the MPI Magdeburg and I'm working in the field of mathematical modeling, model verification, parameter identification, model analysis and experimental design. I'm involved in two projects, KOsmoBac and PSYSMO.
Projects: SysMO-LAB
Institutions: Wageningen University & Research
Expertise: Bioinformatics, Mathematical modelling, Reactor models, dynamics of biological networks.
Tools: Molecular Biology, Cell biology, Computational and theoretical biology, Metabolomics, Model organisms, SBML, ODE, Partial differential equations, Algebraic equations, Linear equations, Copasi, JWS Online, Matlab, Mathematica, SQL, Material balance based modeling
I'm a modeller, specialized in kinetic modeling of biochemical networks. My focus in the SysMO-LAB consortium is on creating models of Lactococcus lactis glycolysis and couple this to other related lactic acid bacteria like Streptococcus pyogenes and Enterococcus faecalis. Besides kinetic modeling, I'm also interested in combining various modeling techniques (genome-scale modeling, qualitative modeling).
Projects: KOSMOBAC
Institutions: University of Aberdeen
Physicist, working on the modelling side.
Projects: COSMIC, BaCell-SysMO, SYSTERACT, HUMET Startup, INCOME, BESTER, iRhythmics, COVID-19 Disease Map, SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke, Implementation of Nanopore Sequencing for Detection of Treatment Induced Transcriptomic and Epitranscriptomic Changes in Leukaemic Tumour Models
Institutions: University of Rostock
Expertise: dynamics of biological networks, Data analysis Mathematical modelling Bioinformatics Systems biology, Dynamics and Control of Biological Networks
Tools: Computational and theoretical biology, ODE, Matlab, Mathematica, stimulus response experiments, differential algebraic equations, quantitative western blot analyses, quantitative western blot analysis, Stochastic models
Projects: COSMIC
Institutions: University of Rostock
biomathematician, PhD student at the University of Rostock, Systems Biology Group Rostock
Expertise: Mathematical modelling, Statistically and biologically inspired optimization algorithms, Mathematical modelling of biosystems and bioprocesses, Dynamics and Control of Biological Networks, Parameter estimation
Tools: Computational and theoretical biology, ODE, Partial differential equations, Matlab, Mathematica, Computational Systems Biology, including: - Dynamic modelling - Parameter estimation - Optimal experimental design - Dynamic optimization
We are very interested in applying a systems approach (i.e. model-based concepts and related computational tools) to problems from the biological domain. In particular, we are doing research in computational systems biology, targetting the following topics:
- Parameter estimation (inverse problems, model calibration) in biochemical pathways
- Optimal experimental design (optimal dynamic experiments) for Systems Biology
- Dynamic optimization (optimal control) of biosystems and bioprocesses
...
Projects: PSYSMO
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: KOSMOBAC
Institutions: University of Aberdeen
My background is physics engineering & biomedical engineering. I did my PhD in Surrey on the modelling of response of mammalian cells to radiation of different qualities. I have been working at the University of Aberdeen since November 2007 as a theoreticien research fellow of the KOSMOBAC project. We are investigating the homeostasis of ions in bacteria E. coli. I have been working at a model of the buffering capacity of the cytopplasm, arising from the presence of weak acids and bases. We ...
Projects: KOSMOBAC
Institutions: University of Aberdeen
Expertise: Mathematical modelling, Mathematical modelling of biosystems and bioprocesses
Tools: Computational and theoretical biology, ODE, linux, data modeling, enzymatic analyses, Stochastic models, C programming, Computational Systems Biology, Deterministic models, including:- Dynamic modelling- Parameter estimation- Optimal experimental design- Dynamic optimization
Projects: KOSMOBAC
Institutions: University of Aberdeen
Expertise: Mathematical modelling, Physics, Programming
Tools: Computational and theoretical biology, ODE, linux, Stochastic models, PDE, C programing
Expertise: Mathematical modelling of biosystems and bioprocesses, Optimal experimental design, Systems Biology, sensitivity analysis, Dynamic optimization., Dynamics and Control of Biological Networks, Parameter estimation
Tools: ODE, Partial differential equations, Matlab, differential algebraic equations, Stochastic models, Computational Systems Biology, Deterministic models, Dynamic modelling, Parameter estimation
I am a postdoctoral researcher in the group of Julio Banga. My research is focused on computational systems biology with particular attention to the mathematical modelling of biosystems and bioprocesses. Some of the topics we address are:
- Parameter estimation
- Model identifiability
- Global sensitivity analysis
- Optimal experimental design
- Dynamic optimization
- Robust control of diffusion-reaction systems
Projects: KOSMOBAC
Institutions: Max Planck Institute for Dynamics of Complex Technical Systems
Projects: BaCell-SysMO
Institutions: University of Rostock
Expertise: Mathematical modelling, Bacillus subtilis, Deterministic modelling of gene regulation networks, stress responses, Systems Biology, sensitivity analysis, Dynamics and Control of Biological Networks, Parameter estimation
Tools: Biochemistry, Computational and theoretical biology, ODE, Matlab, linux, Stochastic models, Deterministic models, Dynamic modelling
Modelling of the general stress response activation cascade of sigB in B. subtilis in response to starvation.
Projects: TRANSLUCENT
Institutions: University of Applied Sciences Koblenz, Rhein Ahr Campus
Grammar school until 1998 1998-99 Alternative civilian service 1999-2002 Professional education as male nurse 2002-2007 Study of Biomathematics 2007-... PhD student in TRANSLUCENT project
Projects: SUMO
Institutions: University of Stuttgart
Expertise: Mathematical modelling, Data Management, Systems Biology, Parameter estimation
Tools: SBML, ODE, Matlab, Mathematica
Former: PhD student as research associate at the Institute for System Dynamics (ISYS), Universität Stuttgart, Germany. Engineering background→modelling, identification and analyses. Detailed kinetic modelling, identification and analysis of the TCA cycle (tricarboxylic acid cycle, citric acid cycle) and the ETC (electron transport chains, respiratory chains) of Escherichia coli. One of the SysMO-DB pals for SUMO. Now: Industrial affiliation
Projects: COSMIC, BaCell-SysMO
Institutions: University of Rostock
Expertise: Mathematical modelling, dynamics of biological networks, Physics, bistability, Systems Biology, Data analysis, Statistical Physics, Dynamics and Control of Biological Networks
Tools: Computational and theoretical biology, ODE, Matlab, Mathematica, differential algebraic equations, Stochastic models, C programming, Computational Systems Biology, Dynamic modelling, Data Modelling
Modelling of cellular signalling, Dynamic Motifs and Feedback, Quantitative Measures, Theoretical Aspects of Modelling Biological Systems
Projects: BaCell-SysMO
Institutions: University of Stuttgart
Expertise: Microbiology, Biochemistry, Mathematical modelling, Bacillus subtilis, Mathematical modelling of biosystems and bioprocesses, stress responses, Systems Biology, Nonlinear Dynamics, carbon metabolism, Signalling networks, Metabolic Networks
Tools: Computational and theoretical biology, ODE, Matlab, Mathematica, Fermentation, Chromatography, continuous cultivation, Enzyme assay, Computational Systems Biology, Deterministic models, Dynamic modelling, fed-batch cultivation
I am a biologist in the lab of Prof. Reuss at the University of Stuttgart and I am working in the field of biotechnology and mathematical modelling.
Projects: SysMO-LAB
Institutions: University of Heidelberg
I am working on a kinetic model of the central metabolism as well as on a genome wide model of Streptococcus pyogenes.
Projects: COSMIC
Institutions: University of Nottingham
Expertise: Mathematical modelling
Tools: Computational and theoretical biology, ODE, Partial differential equations
The dataset presents mathematical models of the gene regulatory network of the circadian clock, in the plant Arabidopsis thaliana. The work will be published as Urquiza-Garcia, Molina, Halliday and Millar, title "Abundant clock proteins point to missing molecular regulation in the plant circadian clock", in Molecular Systems Biology, 2025 doi 10.1038/s44320-025-00086-5.
Starting from the U2019.3 and U2020.3 models, this project rescales parameters to match protein levels that were predicted ...
Submitter: Andrew Millar
Studies: Construction of NanoLUC-tagged plants, Estimating DNA-binding affinities for Arabidopsis proteins, Measuring absolute levels of clock proteins with calibrated NanoLUC assays, Predicting absolute levels of clock proteins with a simple model, Recalibrating the clock models for absolute protein levels, to create mo..., Reproducibility documentation
Assays: Clock protein number determination with NanoLUC calibration, Clock proteins NanoLUC fusion raw data, Gatway maps of genomic regions of clock genes, In vivo bioluminescence of clock protein-NanoLUC fusions: example experi..., Jupyter notebook Predicting Protein Numbers, Propagating scaling factors into model parameters for U2019.4->U2019.5 a..., Protein level time series, Python packages, Reproducibility tool set, Selection of complemented transgenic lines, TiMet RNA timeseries data, promoter binding affinity calculations on the genome based on PBMs and E...
Snapshots: Snapshot 1
The models in this record were published in Flis et al. Royal Society Open Biology 2015. They will be submitted to Biomodels when we have a PubMed ID for the paper.
Original model: Arabidopsis clock model P2011.1.1 from Pokhilko et al. Mol Syst. Biol. 2012, http://dx.doi.org/10.1038/msb.2012.6
Published version is Biomodels ID 00412, http://www.ebi.ac.uk/compneur-srv/biomodels-main/BIOMD0000000412 Also public in Plasmo as PLM_64, with several versions, http://www.plasmo.ed.ac.uk/plasmo/models/model.shtml?accession=PLM_64 ...
Submitter: BioData SynthSys
Biological problem addressed: Gene Regulatory Network
Investigation: Millar, Andrew (ex-PlaSMo models)
Organisms: Arabidopsis thaliana
Models: Arabidopsis clock model P2011.2.1 - PLM_71, ver..., Arabidopsis clock model P2011.2.1 - PLM_71, ver...
SOPs: No SOPs
Data files: SBSI output from optimisation 26-30 July 2013, ..., Unpacked SBSI optimisation results, PLM_71_ 2
Snapshots: No snapshots
RNA timeseries data for Arabidopsis Col wild-type plants and clock mutants, as separate mean and SD files. The raw data is available on BioDare.ed.ac.uk, and is linked as 'Attribution' from elsewhere on FAIRDOMHub.
The starting models are included here in their original forms, the P2011 model as an SBML L3V1 model file, and the KF2014 model of Fogelmark et al. shared as SBML; both prepared by Uriel Urquiza.
Submitter: Andrew Millar
Biological problem addressed: Gene Regulatory Network
Investigation: Absolute units in Arabidopsis clock models up t...
Organisms: No organisms
Models: Arabidopsis clock model P2011.1.2, F2014 all parameters in SBML, F2014.1 - PLM_1030, version 1, SUBMITTED, F2014.1.2 with stepfunction and 1 hidden item
SOPs: No SOPs
Data files: Processed TiMet WP1.1a RNA data, SD, Processed TiMet WP1.1a RNA data, mean
Snapshots: No snapshots
Biochemical Reaction Networks Biochemical Reaction Networks transform static protein-protein interactions (PPI) network into dynamic, mechanistic models by decomposing each interaction into underlying molecular events like phosphorylation and complex formation. Using SPADAN toolbox (DOI:10.1093/bioinformatics/btad079), we expands PPI network into biochemical reactions network.
Ordinary Differential Equation Model Equations Kinetic descriptions of biochemical reactions related to each renal cell ...
Creator: Farnoush Kiyanpour
Submitter: Farnoush Kiyanpour
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
The FEM models how metabolic slowdown will induce the age-related changes of weight gain, insulin resistance, basal inflammation, mitochondrial dysfunction, as well as the age-related disease of atherosclerosis, via a series of unavoidable homeostatic shifts.
Creators: James Wordsworth, Pernille Yde Nielsen
Submitter: James Wordsworth
Model type: Ordinary differential equations (ODE)
Model format: R code
Environment: Not specified
Organism: Not specified
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Model associated with the following:
Hannah A Kinmonth-Schultz, Melissa J S MacEwen, Daniel D Seaton, Andrew J Millar, Takato Imaizumi, Soo-Hyung Kim, An explanatory model of temperature influence on flowering through whole-plant accumulation of FLOWERING LOCUS T in Arabidopsis thaliana, in silico Plants, Volume 1, Issue 1, 2019, diz006, https://doi.org/10.1093/insilicoplants/diz006
Creator: Hannah Kinmonth-Schultz
Submitter: Hannah Kinmonth-Schultz
Model type: Not specified
Model format: Matlab code
Environment: Matlab
Organism: Arabidopsis thaliana
Investigations: Temperature effects on Arabidopsis floral induc...
Originally submitted model file for PLaSMo accession ID PLM_71, version 1
Creators: BioData SynthSys, Andrew Millar, Andrew Millar
Submitter: BioData SynthSys
Model type: Ordinary differential equations (ODE)
Model format: SBML
Environment: Not specified
Organism: Arabidopsis thaliana
Investigations: Millar, Andrew (ex-PlaSMo models)
Originally submitted model file for PLaSMo accession ID PLM_71, version 2
Creators: BioData SynthSys, Andrew Millar, Andrew Millar
Submitter: BioData SynthSys
Model type: Ordinary differential equations (ODE)
Model format: SBML
Environment: Copasi
Organism: Arabidopsis thaliana
Investigations: Millar, Andrew (ex-PlaSMo models)
Here is a kinetic model (in COPASI format) of L. lactis glycolysis.
Creator: Mark Musters
Submitter: Mark Musters
Model type: Ordinary differential equations (ODE)
Model format: Copasi
Environment: Copasi
Organism: Lactococcus lactis
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Abstract (Expand)
Authors: Yin Hoon Chew, Daniel D Seaton, Virginie Mengin, Anna Flis, Sam T Mugford, Gavin M George, Michael Moulin, Alastair Hume, Samuel C Zeeman, Teresa B Fitzpatrick, Alison M Smith, Mark Stitt, Andrew J Millar
Date Published: 1st Jul 2022
Publication Type: Journal Article
DOI: 10.1093/insilicoplants/diac010
Citation: in silico Plants 4(2),diac010
Abstract (Expand)
Authors: Yin Hoon Chew, Daniel D. Seaton, Virginie Mengin, Anna Flis, Sam T. Mugford, Gavin M. George, Michael Moulin, Alastair Hume, Samuel C. Zeeman, Teresa B. Fitzpatrick, Alison M. Smith, Mark Stitt, Andrew J. Millar
Date Published: 6th Feb 2017
Publication Type: Report
DOI: 10.1101/105437
Citation: biorxiv;105437v2,[Preprint]
Abstract (Expand)
Authors: Uriel Urquiza Garcia, Andrew J Millar
Date Published: 5th Aug 2021
Publication Type: Journal Article
DOI: 10.1093/insilicoplants/diab022
Citation:
Abstract (Expand)
Authors: Hannah A Kinmonth-Schultz, Melissa J S MacEwen, Daniel D Seaton, Andrew J Millar, Takato Imaizumi, Soo-Hyung Kim
Date Published: 2019
Publication Type: Journal Article
DOI: 10.1093/insilicoplants/diz006
Citation: in silico Plants 1(1),diz006
Abstract (Expand)
Editor:
Date Published: 11th Nov 2010
Publication Type: Not specified
PubMed ID: 21063964
Citation:
 Download
Download