This is the exchange platform of the COMBINE network.
COMBINE is an initiative to coordinate the development of various community standards and formats for computational models: BioPax, CellML, NeuroML, Synthetic Biology Open Language (SBOL), Systems Biology Graphical Notation (SBGN), Systems Biology Markup Language (SBML), Simulation Experiment Description Markup Language (SED-ML).
Web page: http://co.mbine.org
Funding details:Related items
- People (30)
- Projects (2)
- Institutions (19)
- Investigations (1+1)
- Studies (0+1)
- Assays (0+1)
- Data files (1+2)
- Models (2+1)
- Publications (8)
- Presentations (2+1)
- Events (3)
- Documents (4)
Projects: de.NBI-SysBio, EnzymeML, COMBINE Multicellular Modelling, Standardization of enzyme-catalyzed reaction measurement, Standardization of enzyme-catalyzed reaction modelling
Institutions: University of Heidelberg
 https://orcid.org/0000-0001-5553-4702
https://orcid.org/0000-0001-5553-4702
Lutz Brusch is heading the research group "Spatio-temporal pattern formation in cells and tissues" at the Centre for Information Services and High Performance Computing of TU Dresden, Germany. The group is co-developing the multi-cellular modelling and simulation framework Morpheus (https://morpheus.gitlab.io) and is collaborating with experimental labs on questions of tissue morphogenesis and regeneration.
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0001-8003-6860
https://orcid.org/0000-0001-8003-6860
Expertise: Mathematical modelling, Multicellular Modelling
Research Scientist, Computer Vision; Intellisense Systems, Inc. Lead Developer for MultiCellDS
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0003-3634-190X
https://orcid.org/0000-0003-3634-190X
Expertise: Computational Biology, Mathematical modelling, Multicellular Modelling, Compucell3D, Virtual Tissues, Model Sharing, Language Standards, Agent-based modelling, Dynamic modelling, Computational Systems Biology, standards, Developmental Biology, Toxicology, Framework Development, Cancer, Immunology, Community Building
Tools: Compucell3D, Antimony, tellurium, SBML, CC3DML
Dr. Glazier’s research focuses on early embryonic development, developmental and chronic toxicity and disease, with more than 100 experimental and computational papers on biological development and developmental diseases (including polycystic kidney disease (ADPKD), tumor growth and vascularization, Age Related Macular Degeneration and diabetic retinopathies, somitogenesis and liver toxicity) and more recently on modeling in-host viral infection and immune response. As part of his work on infection ...
Projects: COMBINE Multicellular Modelling
Institutions: University College London (UCL)
 https://orcid.org/0000-0001-5963-8576
https://orcid.org/0000-0001-5963-8576
Expertise: Dynamic modelling, Databases, Mathematical modelling, standards, Neuroscience, NeuroML
Projects: FAIRDOM, Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, MS_DILI, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training, EnzymeML, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", FAIRDOM Community Workers, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, ModeleXchange initiative, SDBV/HITS, EDITH (Ecosystem Digital Twins in Health) test project
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-8683-7084
https://orcid.org/0000-0002-8683-7084
Data management and standardization expert for systems biology and systems medicine, responsible for the data management user requirements and user contacts within the German LiSyM network (Liver Systems Medicine: http://lisym.org/) and associated to the FAIRDOM team. Involved in different standardization initiatives and committees, i.e. COMBINE (http://co.mbine.org), ISO/TC 276 Biotechnology (https://www.iso.org/committee/4514241.html), European COST action CHARME (http://www.cost-charme.eu) and ...
Projects: COMBINE Multicellular Modelling
Institutions: Indiana University Bloomington
 https://orcid.org/0000-0002-7440-2905
https://orcid.org/0000-0002-7440-2905
Expertise: Mathematical modelling, Data Management, Software Engineering, Python, standards, c++
Research Associate in the Macklin Lab, School of Informatics, Computing, and Engineering. Indiana University, Bloomington, IN USA.
Projects: COVID-19 Disease Map, ModeleXchange initiative
Institutions: The European Bioinformatics Institute < EMBL-EBI
Expertise: Knowledge integration, Curation
Projects: FAIRDOM, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, LiSyM Core Infrastructure and Management (LiSyM-PD), ModeleXchange initiative, SDBV/HITS, EDITH (Ecosystem Digital Twins in Health) test project
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-8318-3222
https://orcid.org/0000-0002-8318-3222
Projects: COMBINE Multicellular Modelling
Institutions: University of St Andrews
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
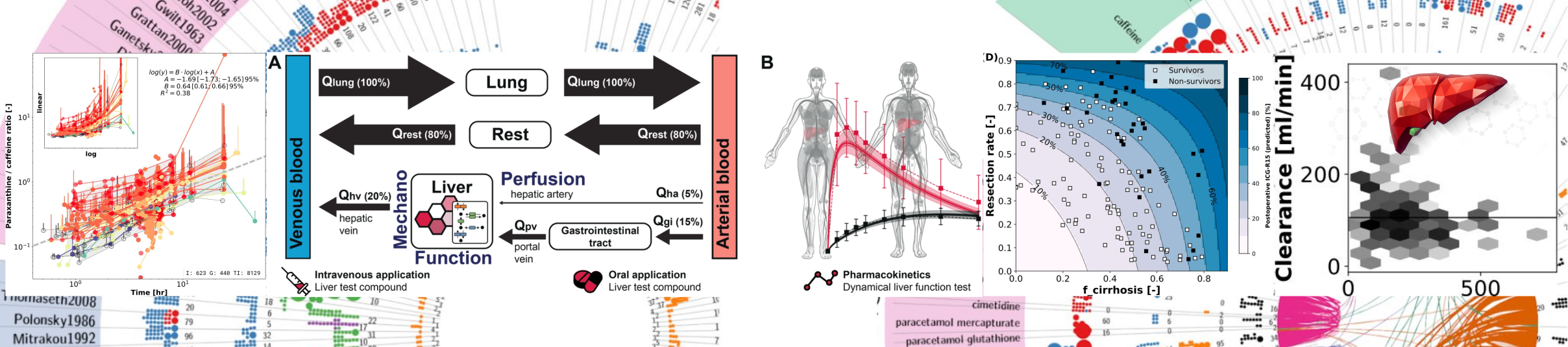
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: COMBINE Multicellular Modelling
Institutions: University of Newcastle-upon-Tyne
 https://orcid.org/0000-0002-8361-2795
https://orcid.org/0000-0002-8361-2795
Interdisciplinary researcher; passionate about findable, accessible, interoperable, and reproducible (FAIR) scientific knowledge.
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, de.NBI-SysBio, Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, EnzymeML, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, MS_DILI, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COMBINE Multicellular Modelling, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, NMTrypI - New Medicines for Trypanosomatidic Infections, ModeleXchange initiative, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, BioCreative VII, SDBV ephemeral data exchanges, The BeeProject, SDBV/HITS, MESI-STRAT Review, MESI-Review 2024, DeepCurate, SABIO-VIS
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-4980-3512
https://orcid.org/0000-0002-4980-3512
I am group leader of the SDBV (Scientific Databases and Visualisation) group at the HITS gGmbH, the Heidelberg Institute for Theoretical Studies.
I am interested in finding data. Starting with my master's thesis I have always worked on how to store data in a way that you can find it, and how to make sense out of data that has been stored.
Within FAIRDOM I find interesting to help people to store their data in a way that they make sense even after years.
Projects: COMBINE Multicellular Modelling
Institutions: University of Heidelberg
 https://orcid.org/0000-0002-7637-7547
https://orcid.org/0000-0002-7637-7547
Programme: COMBINE (Computational Modeling in Biology Network)
Public web page: https://multicellml.org/
Organisms: Not specified
Programme: COMBINE (Computational Modeling in Biology Network)
Public web page: Not specified
Organisms: Not specified
ROR ID: Not specified
Department: Not specified
Country:  New Zealand
New Zealand
City: Auckland
Web page: http://www.abi.auckland.ac.nz/
ROR ID: Not specified
Department: Not specified
Country:  Italy
Italy
City: Milano
Web page: http://bimib.disco.unimib.it/
ROR ID: Not specified
Department: Not specified
Country:  United Kingdom
United Kingdom
City: Hinxton
Web page: https://www.ebi.ac.uk/
ROR ID: https://ror.org/01f7bcy98
Department: Not specified
Country:  Germany
Germany
City: Heidelberg
Web page: https://www.h-its.org
ROR ID: Not specified
Department: Not specified
Country:  Germany
Germany
City: Berlin
Web page: http://www.hu-berlin.de/
ROR ID: Not specified
Department: Not specified
Country:  United States
United States
City: New York
Web page: http://icahn.mssm.edu/
ROR ID: Not specified
Department: Not specified
Country:  United States
United States
City: Bloomington
Web page: https://www.indiana.edu
ROR ID: Not specified
Department: Not specified
Country:  Germany
Germany
City: 44139 Dortmund
Web page: http://www.ifado.de/
ROR ID: Not specified
Department: Not specified
Country:  Norway
Norway
City: Trondheim
Web page: http://www.ntnu.edu/
ROR ID: Not specified
Department: Not specified
Country:  South Africa
South Africa
City: Not specified
Web page: Not specified
ROR ID: Not specified
Department: Not specified
Country:  Germany
Germany
City: Dresden
Web page: http://tu-dresden.de/en
ROR ID: Not specified
Department: Not specified
Country:  United Kingdom
United Kingdom
City: Hinxton, Cambridgeshire
Web page: Not specified
ROR ID: Not specified
Department: Not specified
Country:  United Kingdom
United Kingdom
City: London
Web page: https://www.ucl.ac.uk
ROR ID: Not specified
Department: Not specified
Country:  Netherlands
Netherlands
City: Maastricht
Web page: Not specified
ROR ID: Not specified
Department: Not specified
Country:  United States
United States
City: Not specified
Web page: Not specified
ROR ID: Not specified
Department: Not specified
Country:  United Kingdom
United Kingdom
City: Not specified
Web page: Not specified
ROR ID: Not specified
Department: Not specified
Country:  Germany
Germany
City: Rostock
Web page: http://www.uni-rostock.de
ROR ID: Not specified
Department: Not specified
Country:  United Kingdom
United Kingdom
City: St Andrews
Web page: https://www.st-andrews.ac.uk
Virtual Tissues (VTs) are multi-scale, multi-cellular, mechanistic Agent Based Models (ABMs) that predict the spatio-temporal dynamics of biological tissues. While multiple platforms exist for constructing and executing VT models (listed below in the section on VT Modeling Frameworks), models developed for different platforms are currently incompatible and not accessible or executable in a common location, impeding model discovery, validation and reuse. FAIRSPACE will initially provide support ...
Snapshots: No snapshots
This mini-symposium, colocated at the ECMTB in Gothenburg, presents the state-of-the-art in computational methods for multicellular systems biology. It brings together developers and users of this software to identify common approaches and future challenges concerning multiscale integration, model specification, model exchange, scalability, workflow management as well as compliance to standards and guidelines. The session thus aims to provide an overview of the available modeling and simulation ...
Creator: Walter de Back, TU Dresden
Submitter: Lutz Brusch
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
For the spatio-temporal dynamics of bile transport, bile canalicular dilation, mechanical stimulation and transduction of YAP signaling during liver regeneration see the open access publication and its appendix: Meyer et al. (2020) Bile canaliculi remodeling activates YAP via the actin cytoskeleton during liver regeneration. Molecular Systems Biology 16:e8985. https://doi.org/10.15252/msb.20198985
The model format is MorpheusML that can readily be loaded and run in the free and open source software ...
Creator: Lutz Brusch
Submitter: Lutz Brusch
Model type: Ordinary differential equations (ODE)
Model format: Morpheus
Environment: Morpheus
Spatio-temporal liver zonation in mouse and human with Wnt-Hh crosstalk and transport are modeled using coupled partial differential equations. The model file is in MorpheusML format and can be opened in the free, open-source multicellular modeling software Morpheus (https://morpheus.gitlab.io). In Morpheus, the model will simulate the time course (movie) of dynamic liver zonation for a 2D cross-section of several liver lobules, showing the patterns of Wnt ligands, intracellular Wnt signaling, ...
Creators: Lutz Brusch, Jörn Starruß, Michael Kücken
Submitter: Lutz Brusch
Model type: Partial differential equations (PDE)
Model format: Morpheus
Environment: Morpheus
Abstract (Expand)
Authors: Dagmar Waltemath, Martin Golebiewski, Michael L Blinov, Padraig Gleeson, Henning Hermjakob, Michael Hucka, Esther Thea Inau, Sarah M Keating, Matthias König, Olga Krebs, Rahuman S Malik-Sheriff, David Nickerson, Ernst Oberortner, Herbert M Sauro, Falk Schreiber, Lucian Smith, Melanie I Stefan, Ulrike Wittig, Chris J Myers
Date Published: 29th Jun 2020
Publication Type: Journal Article
Citation: Journal of Integrative Bioinformatics 0(0)
Abstract (Expand)
Authors: Falk Schreiber, Björn Sommer, Tobias Czauderna, Martin Golebiewski, Thomas E. Gorochowski, Michael Hucka, Sarah M. Keating, Matthias König, Chris Myers, David Nickerson, Dagmar Waltemath
Date Published: 29th Jun 2020
Publication Type: Journal Article
Citation: Journal of Integrative Bioinformatics 0(0)
Abstract (Expand)
Authors: N. J. Stanford, M. Scharm, P. D. Dobson, M. Golebiewski, M. Hucka, V. B. Kothamachu, D. Nickerson, S. Owen, J. Pahle, U. Wittig, D. Waltemath, C. Goble, P. Mendes, J. Snoep
Date Published: 12th Oct 2019
Publication Type: Journal Article
PubMed ID: 31602618
Citation: Methods Mol Biol. 2019;2049:285-314. doi: 10.1007/978-1-4939-9736-7_17.
Abstract (Expand)
Authors: Falk Schreiber, Björn Sommer, Gary D. Bader, Padraig Gleeson, Martin Golebiewski, Michael Hucka, Sarah M. Keating, Matthias König, Chris Myers, David Nickerson, Dagmar Waltemath
Date Published: 13th Jul 2019
Publication Type: Journal Article
Citation: Journal of Integrative Bioinformatics 16(2)
Abstract (Expand)
Authors: Maxwell Lewis Neal, Matthias König, David Nickerson, Göksel Mısırlı, Reza Kalbasi, Andreas Dräger, Koray Atalag, Vijayalakshmi Chelliah, Michael T Cooling, Daniel L Cook, Sharon Crook, Miguel de Alba, Samuel H Friedman, Alan Garny, John H Gennari, Padraig Gleeson, Martin Golebiewski, Michael Hucka, Nick Juty, Chris Myers, Brett G Olivier, Herbert M Sauro, Martin Scharm, Jacky L Snoep, Vasundra Touré, Anil Wipat, Olaf Wolkenhauer, Dagmar Waltemath
Date Published: 1st Mar 2019
Publication Type: Journal Article
DOI: 10.1093/bib/bby087
Citation: Briefings in Bioinformatics 20(2):540-550
Abstract (Expand)
Author: Martin Golebiewski
Date Published: 2019
Publication Type: Book Chapter
DOI: 10.1016/B978-0-12-809633-8.20471-8
Citation: Encyclopedia of Bioinformatics and Computational Biology,pp.884-893,Elsevier
Abstract (Expand)
Authors: Chris J. Myers, Gary Bader, Padraig Gleeson, Martin Golebiewski, Michael Hucka, Nicolas Le Novere, David P. Nickerson, Falk Schreiber, Dagmar Waltemath
Date Published: 1st Dec 2017
Publication Type: Conference Paper
Citation: 2017 Winter Simulation Conference (WSC),pp.884-895,IEEE
Abstract (Expand)
Authors: K. Wolstencroft, O. Krebs, J. L. Snoep, N. J. Stanford, F. Bacall, M. Golebiewski, R. Kuzyakiv, Q. Nguyen, S. Owen, S. Soiland-Reyes, J. Straszewski, D. D. van Niekerk, A. R. Williams, L. Malmstrom, B. Rinn, W. Muller, C. Goble
Date Published: 4th Jan 2017
Publication Type: Journal Article
PubMed ID: 27899646
Citation: Nucleic Acids Res. 2017 Jan 4;45(D1):D404-D407. doi: 10.1093/nar/gkw1032. Epub 2016 Nov 28.
An overview over important lessons from the experience with SBML Level 3 extension for the MultiCellML language.
Creator: Matthias König
Submitter: Matthias König
The “Computational Modeling in Biology” Network (COMBINE) is an initiative to coordinate the development of the various community standards and formats in systems biology and related fields. COMBINE 2022 will be a workshop-style event with oral presentations, breakout sessions and tutorials. The three meeting days will include talks about the COMBINE standards and associated or related standardization efforts, presentations of tools using these standards, breakout sessions for detailed discussions ...
Start Date: 6th Oct 2022 (6th Oct 2022 (Etc/UTC))
End Date: 8th Oct 2022 (8th Oct 2022 (Etc/UTC))
Event Website: https://co.mbine.org/author/combine-2022/
Country:  Germany
Germany
City: Berlin
Aim Multicellular spatial models have become an essential tool in quantitative biology and multiple model formalisms and simulation frameworks coexist. Progress towards software interoperability and reproducibility requires a novel model description language and conversion library, complementing SBML. Building on the community‘s practical experiences with multiple formalisms and simulators, we aim at (1) defining the concept for a multicellular language while simultaneously (2) exploring automated ...
Start Date: 1st Sep 2019
End Date: 4th Sep 2019
Event Website: https://multicellml.org/wiki/doku.php?id=hackathon
Country:  Germany
Germany
City: Dresden
This mini-symposium, colocated at the ECMTB in Gothenburg, presents the state-of-the-art in computational methods for multicellular systems biology. It brings together developers and users of this software to identify common approaches and future challenges concerning multiscale integration, model specification, model exchange, scalability, workflow management as well as compliance to standards and guidelines. The session thus aims to provide an overview of the available modeling and simulation ...
Start Date: 19th Jun 2014
End Date: 19th Jun 2014
Event Website: https://imc.zih.tu-dresden.de/wiki/multisysbio2014/
Country:  Sweden
Sweden
City: Gothenburg
Program of COMBINE sessions on 6th, 7th Oct. 2022 and COMBINE Forum on 8th Oct. 2022.
Creator: Lutz Brusch
Submitter: Lutz Brusch
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Creators: Lutz Brusch, Jörn Starruß
Submitter: Lutz Brusch
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Creators: Lutz Brusch, Jochen Kursawe, Jörn Starruß
Submitter: Lutz Brusch
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Creators: Juliano Ferrari Gianlupi, Lutz Brusch, Jörn Starruß
Submitter: Juliano Ferrari Gianlupi
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
 Download
Download