People
Currently I focuse on the integration of data into multi-scale models with statistical methods and uncertainty tracking in the research unit QuaLiPerF.
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: MIX-UP
Institutions: RWTH Aachen University
Projects: SysMO-LAB, de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases
Institutions: Heidelberg Institute for Theoretical Studies
I'm a biologist working in the field of scientific datases as a biocurator.
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
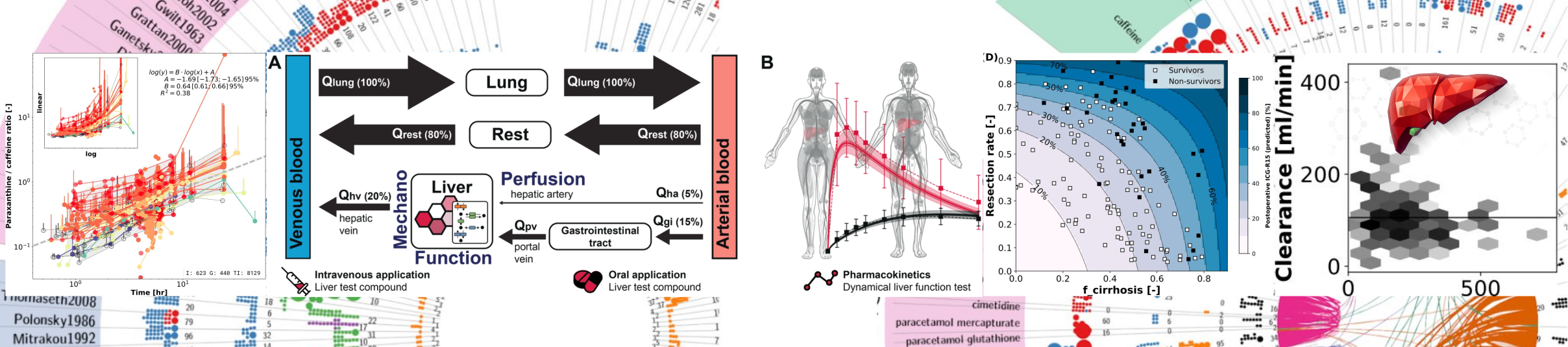
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, ZucAt, SysMO-LAB, Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, ErasysApp Funders, EraCoBiotech 2 nd call proposal preparation, Service to URV Tarragona, Spain with respect to their Safety Assessment of Endocrine Disrupting Chemicals model (Active NOW), FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, Multiscale modelling of state transitions in the host-microbiome-brain network, BESTER, TRALAMINOL, Sustainable co-production, INDIE - Biotechnological production of sustainable indole, Extremophiles metabolsim, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, OLCIR: Optimization of Lung Cancer Therapy with Ionizing Radiation, NAD COMPARTMENTATION, HOTSOLUTE, Stress granules, FAIRDOM Community Workers, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", Mechanism based modeling viral disease ( COVID-19 ) dynamics in human population, COVID-19 Disease Map, AquaHealth (ERA-BlueBio), LiSyM Core Infrastructure and Management (LiSyM-PD), Early Metabolic Injury (LiSyM-EMI - Pillar I), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Modelling COVID-19 epidemics, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, SCyCode The Autotrophy-Heterotrophy Switch in Cyanobacteria: Coherent Decision-Making at Multiple Regulatory Layers, SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke, CC-TOP, BioCreative VII, MESI-STRAT Review, SDBV/HITS, MESI-Review 2024
Institutions: Heidelberg Institute for Theoretical Studies, FAIRDOM User meeting, Norwegian University of Science and Technology, University of Rostock, University of Innsbruck
 https://orcid.org/0000-0003-3540-0402
https://orcid.org/0000-0003-3540-0402
I am a researcher at the Scientific Databases and Visualization Group at Heidelberg Institute for Theoretical Studies (HITS) , one of the developers of SabioRK - System for the Analysis of Biochemical Pathways - Reaction Kinetics (http://sabiork.h-its.org/) . I am working on design and maintenance of the information systems to store, query and analyse systems biology data; definition and implementation of methods for the integration of data from multiple sources. In **[SySMO-DB ...
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: SysMO-LAB, Kinetics on the move - Workshop 2016, de.NBI-SysBio
Institutions: University of Heidelberg
Ursula Kummer is heading the dept. "Modeling of Biological Processes" at the University of Heidelberg.
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Kinetics on the move - Workshop 2016
Institutions: Heidelberg Institute for Theoretical Studies
Research associate at HITS, software developer
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: ZucAt
Institutions: Novosibirsk State University
I got my Master’s Degree in Biomathematics, Bioinformatics, and Computational Biology 2007 at Novosibirsk State University (NSU). I am working at Computer Proteomics Laboratory, Institute of Cytology and Genetics SB RAS. We develope computer system to analyze the coding features of functional sites by taking into account the exon structure of the gene, to detect the exons involved in shuffling in protein evolution, also to design protein-engineering experiments. Tools developed - SitEx http://www-bionet ...
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
 https://orcid.org/0000-0001-6142-5270
https://orcid.org/0000-0001-6142-5270
Projects: Working Group Nicole Radde
Institutions: University of Stuttgart
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, de.NBI-SysBio, Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, EnzymeML, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, MS_DILI, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COMBINE Multicellular Modelling, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, NMTrypI - New Medicines for Trypanosomatidic Infections, ModeleXchange initiative, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, BioCreative VII, SDBV ephemeral data exchanges, The BeeProject, SDBV/HITS, MESI-STRAT Review, MESI-Review 2024, DeepCurate, SABIO-VIS
Institutions: Heidelberg Institute for Theoretical Studies
 https://orcid.org/0000-0002-4980-3512
https://orcid.org/0000-0002-4980-3512
I am group leader of the SDBV (Scientific Databases and Visualisation) group at the HITS gGmbH, the Heidelberg Institute for Theoretical Studies.
I am interested in finding data. Starting with my master's thesis I have always worked on how to store data in a way that you can find it, and how to make sense out of data that has been stored.
Within FAIRDOM I find interesting to help people to store their data in a way that they make sense even after years.
Software developer for FAIRDOM
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: ZucAt
Institutions: Institute of Cytology and Genetics, Novosibirsk State University
I am Senior Scientist at Institute of Cytology and Genetics SB RAS in Novosibirsk. My research focus is computational genomics: high-throughput sequencing, ChIP-seq, human genome research (HUGO member), high performance computing in bioinformatics
Scientific Project Manager at Luxembourg Centre for Systems Biomedicine, University of Luxembourg http://lcsb.uni.lu