Personalized liver function tests: A Multiscale Computational Model Predicts Individual Human Liver Function From Single-Cell Metabolism
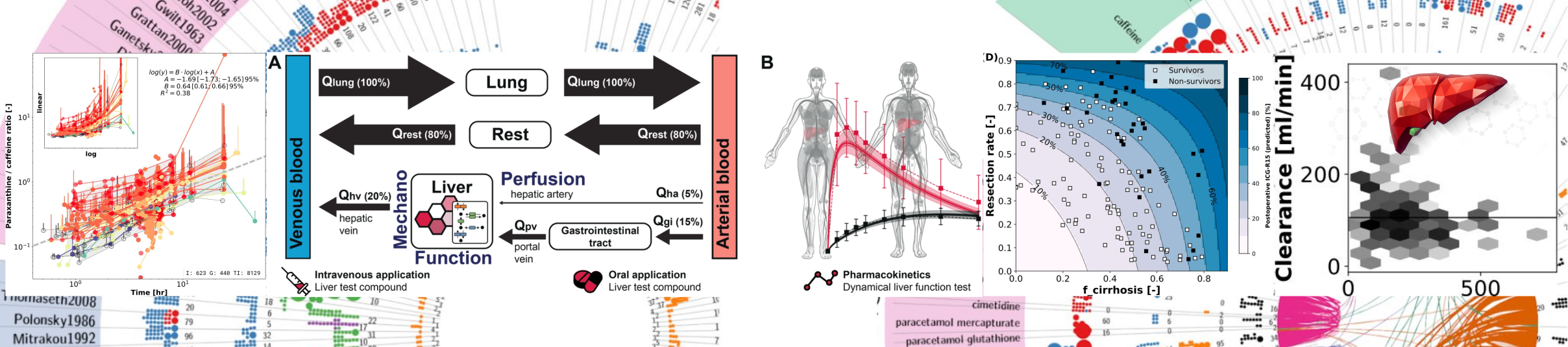
Understanding how liver function arises from the complex interaction of morphology, perfusion, and metabolism from single cells up to the entire organ requires systems-levels computational approaches. We report a multiscale mathematical model of the Human liver comprising the scales from single hepatocytes, over representation of ultra-structure and micro-circulation in the hepatic tissue, up to the entire organ integrated with perfusion. The model was validated against data on multiple spatial and temporal scales. Herein we describe the model construction and application to hepatic galactose metabolism demonstrating its utility via i) the personalization of liver function tests based on galactose elimination capacity (GEC), ii) the explanation of changes in liver function with aging, and iii) the prediction of population variability in liver function based on variability in liver volume and perfusion. We conclude that physiology- and morphology-based multiscale models can improve the evaluation of individual liver function.
Availability: Personalized prediction of liver function (GEC) was implemented in a proof-of-principle app available at https://www.livermetabolism.com/gec_app/. Source code and additional information at https://github.com/matthiaskoenig/multiscale-galactose
 Creators and Submitter
Creators and SubmitterCreator
Submitter
Views: 4350
Created: 16th Aug 2016 at 14:07
Last updated: 3rd May 2017 at 11:24
 Tags
TagsThis item has not yet been tagged.
 Actions
Actions  Order Assays
Order Assays  Download
Download https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X