Tool and work flow development for computational biology.
Investigation position:
Export PNG
 Creators and Submitter
Creators and SubmitterViews: 3026
Created: 25th Jul 2016 at 06:42
 Tags
TagsThis item has not yet been tagged.
Related items
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
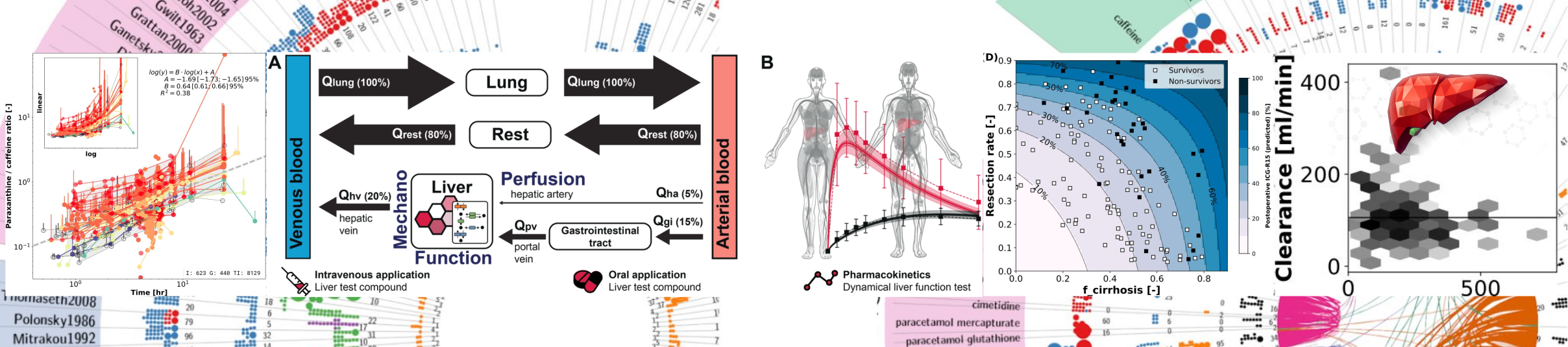
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
LiSyM (Liver Systems Medicine) represents a research network of German centers and institutions, brought together by a 20 Million Euro funding program of the German Government, in which mathematicians, modelers, pharmacologists, molecular biologists and clinical scientists work together to develop a Systems Medicine approach to study early and advanced liver disease. The aim of this unique research program is to acquire and use new experimental data and data from existing data bases to build ...
Projects: Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), FAIRDOM & LiSyM & de.NBI Data Structuring Training, New LiSyM project
Web page: http://www.lisym.org
We will contribute to the LiSyM Research Network an open source, freely available and reproducible multiscale model of the human liver from single cell metabolism to whole liver function. The model will be available in existing standards of systems biology, provide standardized interfaces for data integration and be fully annotated to available biological, medical and computational ontologies. All data, models and source code will be shared within the LiSyM Research Network and made available to ...
Programme: LiSyM - Systems Medicine of the Liver
Public web page: https://livermetabolism.com
Organisms: Homo sapiens, Mus musculus, Rattus norvegicus
Abstract (Expand)
Authors: D. Waltemath, J. Karr, F. Bergmann, V. Chelliah, M. Hucka, M. Krantz, W. Liebermeister, P. Mendes, C. Myers, P. Pir, B. Alaybeyoglu, N. Aranganathan, K. Baghalian, A. Bittig, P. Burke, M. Cantarelli, Y. Chew, R. Costa, J. Cursons, T. Czauderna, A. Goldberg, H. Gomez, J. Hahn, T. Hameri, D. Gardiol, D. Kazakiewicz, I. Kiselev, V. Knight-Schrijver, C. Knupfer, M. Konig, D. Lee, A. Lloret-Villas, N. Mandrik, J. Medley, B. Moreau, H. Naderi-Meshkin, S. Palaniappan, D. Priego-Espinosa, M. Scharm, M. Sharma, K. Smallbone, N. Stanford, J. H. Song, T. Theile, M. Tokic, N. Tomar, V. Toure, J. Uhlendorf, T. Varusai, L. Watanabe, F. Wendland, M. Wolfien, J. Yurkovich, Y. Zhu, A. Zardilis, A. Zhukova, F. Schreiber
Date Published: 16th Jun 2016
Publication Type: Not specified
PubMed ID: 27305665
Citation: IEEE Trans Biomed Eng. 2016 Jun 10.
Abstract (Expand)
Author: Matthias König
Date Published: 2016
Publication Type: Not specified
DOI: 10.12688/f1000research.9211.1
Citation: F1000Res 5 : 1736
 Actions
Actions  Order Studies
Order Studies  Download
Download